Visualize Shape Variation#
Display reconstructed shapes along component axes.
EFA outlines#
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from ktch.datasets import load_outline_mosquito_wings
from ktch.harmonic import EllipticFourierAnalysis
from ktch.plot import shape_variation_plot
data = load_outline_mosquito_wings(as_frame=True)
coords = data.coords.to_numpy().reshape(-1, 100, 2)
efa = EllipticFourierAnalysis(n_harmonics=20)
coef = efa.fit_transform(coords)
pca = PCA(n_components=5).fit(coef)
fig = shape_variation_plot(
pca,
descriptor=efa,
components=(0, 1, 2),
sd_values=(-2, -1, 0, 1, 2),
)
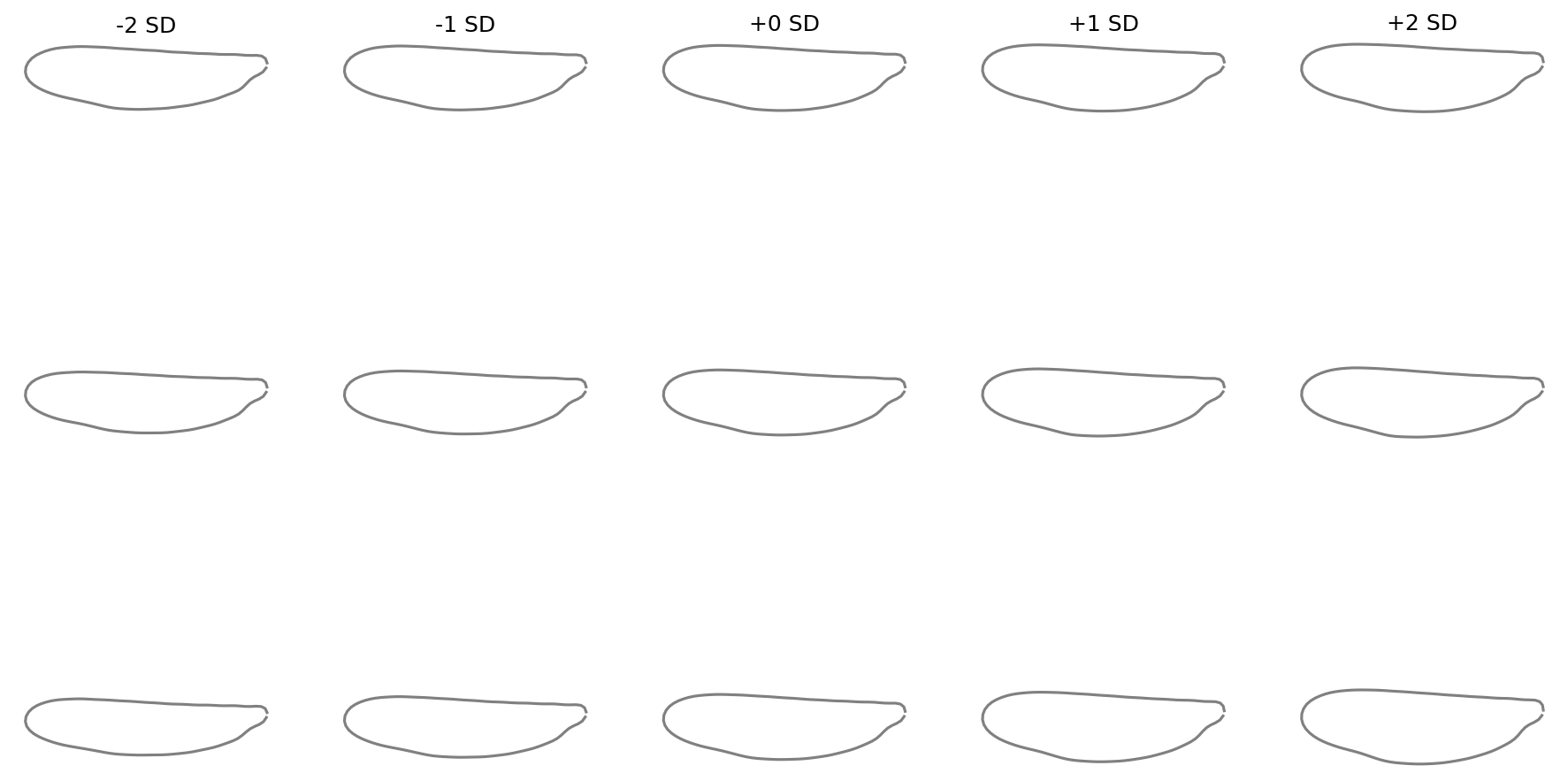
Landmarks with links#
import pandas as pd
from ktch.datasets import load_landmark_mosquito_wings
from ktch.landmark import GeneralizedProcrustesAnalysis
data_lm = load_landmark_mosquito_wings(as_frame=True)
df_coords = (
data_lm.coords.unstack()
.swaplevel(1, 0, axis=1)
.sort_index(axis=1)
)
df_coords.columns = [
dim + "_" + str(idx) for idx, dim in df_coords.columns
]
gpa = GeneralizedProcrustesAnalysis().set_output(transform="pandas")
df_shapes = gpa.fit_transform(df_coords)
# set_output is not needed here: shape_variation_plot only calls
# inverse_transform(), whose return type is unaffected by set_output.
pca_lm = PCA(n_components=5).fit(df_shapes)
links = [
[0, 1], [1, 2], [2, 3], [3, 4], [4, 5], [5, 6],
[6, 7], [7, 8], [8, 9], [9, 10], [10, 11],
[1, 12], [2, 12], [13, 14], [14, 3], [14, 4],
[15, 5], [16, 6], [16, 7], [17, 8], [17, 9],
]
fig = shape_variation_plot(
pca_lm,
n_dim=2,
links=links,
components=(0, 1, 2),
sd_values=(-2, -1, 0, 1, 2),
)
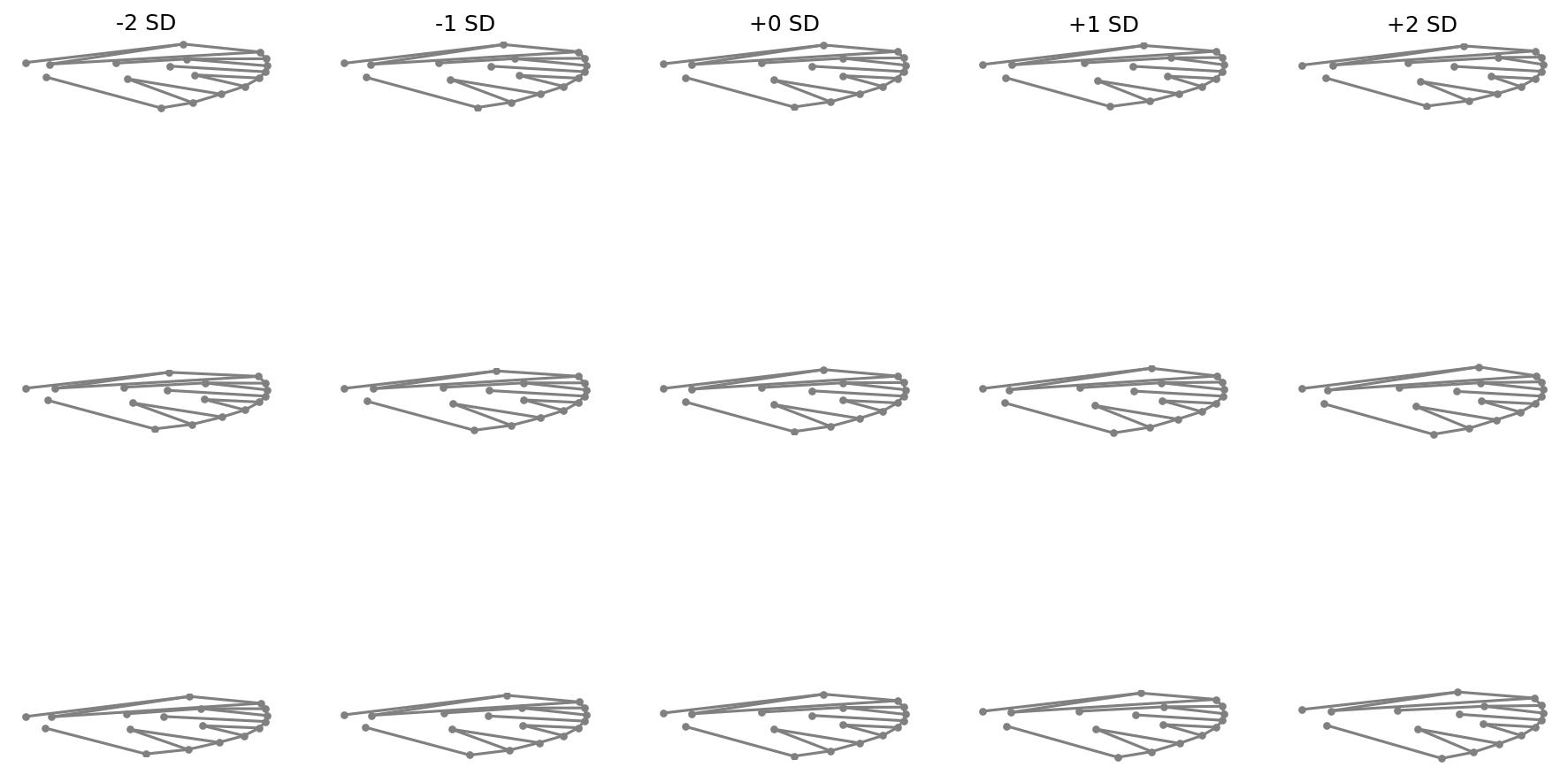
Select components and SD values#
Show only specific components with custom SD steps:
fig = shape_variation_plot(
pca,
descriptor=efa,
components=(0, 1),
sd_values=(-3, -1.5, 0, 1.5, 3),
)
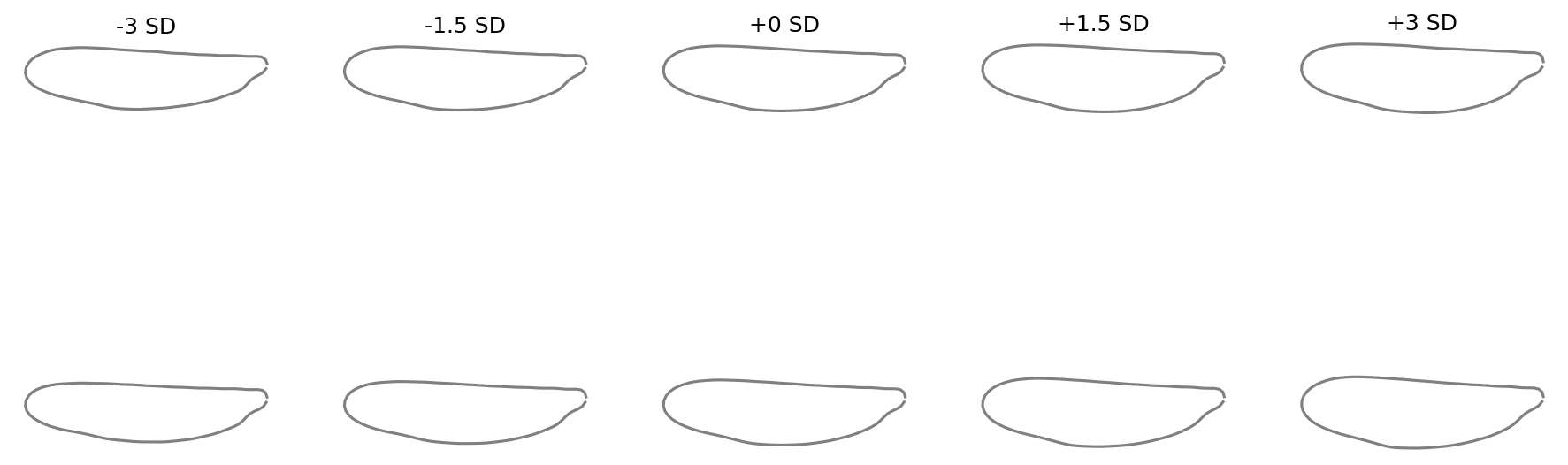
See also
Visualize Morphospace for morphospace scatter plots with shape overlays
Use Non-PCA Reducers for using KernelPCA and other non-PCA reducers
Morphometric Visualization for the reconstruction pipeline design