3D Elliptic Fourier Analysis#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px
from sklearn.decomposition import PCA
from ktch.datasets import load_outline_leaf_bending
from ktch.harmonic import EllipticFourierAnalysis
from ktch.plot import explained_variance_ratio_plot, morphospace_plot
Load the leaf bending outline dataset#
data = load_outline_leaf_bending(as_frame=True)
data.coords
| x | y | z | ||
|---|---|---|---|---|
| specimen_id | coord_id | |||
| 0 | 0 | 0.000000 | 0.000000 | 0.000000 |
| 1 | 0.000000 | -0.046295 | 0.000000 | |
| 2 | 0.000000 | -0.092589 | 0.000000 | |
| 3 | 0.000630 | -0.095962 | 0.045238 | |
| 4 | 0.002576 | -0.098305 | 0.091431 | |
| ... | ... | ... | ... | ... |
| 59 | 195 | 0.000000 | 0.148530 | 0.000000 |
| 196 | 0.000000 | 0.118824 | 0.000000 | |
| 197 | 0.000000 | 0.089118 | 0.000000 | |
| 198 | 0.000000 | 0.059412 | 0.000000 | |
| 199 | 0.000000 | 0.029706 | 0.000000 |
12000 rows × 3 columns
df_meta = data.meta.copy()
df_meta["bending_angle"] = (
np.degrees(df_meta["alphaB"]).round().astype(int).astype(str) + "\u00b0"
)
df_meta["aspect_ratio"] = df_meta["alpha"]
df_meta
| lmax | c | alpha | A | alphaB | kB | alpha_random | alphaB_random | bending_angle | aspect_ratio | |
|---|---|---|---|---|---|---|---|---|---|---|
| specimen_id | ||||||||||
| 0 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.060572 | 2.528513 | 140° | 0.08 |
| 1 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.090911 | 2.318855 | 140° | 0.08 |
| 2 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.088957 | 2.417189 | 140° | 0.08 |
| 3 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.074596 | 2.506030 | 140° | 0.08 |
| 4 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.098990 | 2.294188 | 140° | 0.08 |
| 5 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.073196 | 2.492090 | 140° | 0.08 |
| 6 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.077120 | 2.287193 | 140° | 0.08 |
| 7 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.083971 | 2.280600 | 140° | 0.08 |
| 8 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.073029 | 2.431389 | 140° | 0.08 |
| 9 | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.099243 | 2.553002 | 140° | 0.08 |
| 10 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.089302 | 1.409695 | 80° | 0.08 |
| 11 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.076330 | 1.242604 | 80° | 0.08 |
| 12 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.060501 | 1.299788 | 80° | 0.08 |
| 13 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.087227 | 1.536335 | 80° | 0.08 |
| 14 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.098797 | 1.548685 | 80° | 0.08 |
| 15 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.076723 | 1.280753 | 80° | 0.08 |
| 16 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.093675 | 1.558039 | 80° | 0.08 |
| 17 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.095911 | 1.566630 | 80° | 0.08 |
| 18 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.078236 | 1.387295 | 80° | 0.08 |
| 19 | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.095689 | 1.447270 | 80° | 0.08 |
| 20 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.082873 | 0.390205 | 20° | 0.08 |
| 21 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.064031 | 0.266366 | 20° | 0.08 |
| 22 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.089630 | 0.373168 | 20° | 0.08 |
| 23 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.078860 | 0.330521 | 20° | 0.08 |
| 24 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.087673 | 0.318159 | 20° | 0.08 |
| 25 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.088925 | 0.305914 | 20° | 0.08 |
| 26 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.077160 | 0.372707 | 20° | 0.08 |
| 27 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.083355 | 0.268017 | 20° | 0.08 |
| 28 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.083693 | 0.406805 | 20° | 0.08 |
| 29 | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.081946 | 0.311726 | 20° | 0.08 |
| 30 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.173997 | 2.326146 | 140° | 0.16 |
| 31 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.178168 | 2.367912 | 140° | 0.16 |
| 32 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.166143 | 2.497124 | 140° | 0.16 |
| 33 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.142324 | 2.521001 | 140° | 0.16 |
| 34 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.159544 | 2.492420 | 140° | 0.16 |
| 35 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.176739 | 2.296786 | 140° | 0.16 |
| 36 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.170455 | 2.304695 | 140° | 0.16 |
| 37 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.148710 | 2.478740 | 140° | 0.16 |
| 38 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.141903 | 2.555287 | 140° | 0.16 |
| 39 | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.171313 | 2.610918 | 140° | 0.16 |
| 40 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.176521 | 1.244129 | 80° | 0.16 |
| 41 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.167533 | 1.252522 | 80° | 0.16 |
| 42 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.175097 | 1.438110 | 80° | 0.16 |
| 43 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.146855 | 1.570428 | 80° | 0.16 |
| 44 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.143005 | 1.422087 | 80° | 0.16 |
| 45 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.174511 | 1.497144 | 80° | 0.16 |
| 46 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.170496 | 1.470243 | 80° | 0.16 |
| 47 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.151212 | 1.282264 | 80° | 0.16 |
| 48 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.176779 | 1.257857 | 80° | 0.16 |
| 49 | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.157964 | 1.505475 | 80° | 0.16 |
| 50 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.166466 | 0.346456 | 20° | 0.16 |
| 51 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.177635 | 0.386719 | 20° | 0.16 |
| 52 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.155776 | 0.441191 | 20° | 0.16 |
| 53 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.175787 | 0.407493 | 20° | 0.16 |
| 54 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.146865 | 0.504861 | 20° | 0.16 |
| 55 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.163440 | 0.445301 | 20° | 0.16 |
| 56 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.160243 | 0.366476 | 20° | 0.16 |
| 57 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.158335 | 0.354346 | 20° | 0.16 |
| 58 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.148304 | 0.235357 | 20° | 0.16 |
| 59 | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.160054 | 0.206858 | 20° | 0.16 |
coords = data.coords.to_numpy().reshape(-1, 200, 3)
coords.shape
(60, 200, 3)
Visualize 3D outlines#
representative_ids = [0, 10, 20, 30, 40, 50]
bending_order = [ str(deg)+ "\u00b0" for deg in [20, 80, 140] ]
dfs = []
for sid in representative_ids:
df = pd.DataFrame(coords[sid], columns=["x", "y", "z"])
df["idx"] = data.meta.index[sid]
df["bending_angle"] = df_meta.iloc[sid]["bending_angle"]
dfs.append(df)
df_vis = pd.concat(dfs, ignore_index=True)
fig = px.line_3d(
df_vis,
x="x",
y="y",
z="z",
line_group="idx",
color="bending_angle",
category_orders ={"bending_angle":bending_order},
color_discrete_sequence=px.colors.qualitative.Set2,
)
fig.update_layout(scene=dict(aspectmode="data"))
fig.show()
3D EFA without normalization#
efa = EllipticFourierAnalysis(n_harmonics=20, n_dim=3, norm=False)
coef = efa.fit_transform(coords)
coef.shape
(60, 126)
Reconstruction from coefficients#
coords_recon = efa.inverse_transform(coef, t_num=200)
sid = 0
df_orig = pd.DataFrame(coords[sid], columns=["x", "y", "z"])
df_orig["type"] = "original"
df_rec = pd.DataFrame(coords_recon[sid], columns=["x", "y", "z"])
df_rec["type"] = "reconstructed"
df_cmp = pd.concat([df_orig, df_rec], ignore_index=True)
fig = px.line_3d(df_cmp, x="x", y="y", z="z", color="type")
fig.update_layout(scene=dict(aspectmode="data"))
fig.show()
3D EFA#
efa = EllipticFourierAnalysis(n_harmonics=20, n_dim=3, norm=True)
coef = efa.fit_transform(coords)
coef.shape
(60, 126)
Reconstruction from coefficients#
coords_recon = efa.inverse_transform(coef, t_num=200)
representative_ids = [0, 10, 20, 30, 40, 50]
dfs = []
for sid in representative_ids:
df = pd.DataFrame(coords_recon[sid], columns=["x", "y", "z"])
df["idx"] = data.meta.index[sid]
df["bending_angle"] = df_meta.iloc[sid]["bending_angle"]
dfs.append(df)
df_vis = pd.concat(dfs, ignore_index=True)
fig = px.line_3d(
df_vis, x="x", y="y", z="z",
line_group="idx",
color="bending_angle",
category_orders ={"bending_angle":bending_order},
color_discrete_sequence=px.colors.qualitative.Set2
)
fig.update_layout(scene=dict(aspectmode="data"))
fig.show()
PCA#
pca = PCA(n_components=12)
pcscores = pca.fit_transform(coef)
df_pca = pd.DataFrame(pcscores, columns=[f"PC{i+1}" for i in range(12)])
df_pca.index = df_meta.index
df_pca = df_pca.join(df_meta)
df_pca
| PC1 | PC2 | PC3 | PC4 | PC5 | PC6 | PC7 | PC8 | PC9 | PC10 | ... | lmax | c | alpha | A | alphaB | kB | alpha_random | alphaB_random | bending_angle | aspect_ratio | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| specimen_id | |||||||||||||||||||||
| 0 | -1.212191 | 1.121579 | 0.107171 | 0.011560 | 0.012842 | 0.001138 | -0.002117 | -0.000665 | 5.195792e-04 | 0.000332 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.060572 | 2.528513 | 140° | 0.08 |
| 1 | -0.936970 | 0.436713 | 0.028150 | 0.005296 | -0.004628 | 0.001218 | -0.000426 | -0.000119 | -3.270928e-04 | -0.000379 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.090911 | 2.318855 | 140° | 0.08 |
| 2 | -1.035797 | 0.439970 | 0.047183 | 0.004650 | -0.004887 | 0.000129 | -0.000774 | -0.000030 | -2.139763e-04 | -0.000394 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.088957 | 2.417189 | 140° | 0.08 |
| 3 | -1.145406 | 0.728617 | 0.075460 | 0.008020 | 0.000725 | -0.001096 | -0.000554 | 0.000422 | 2.015761e-04 | -0.000002 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.074596 | 2.506030 | 140° | 0.08 |
| 4 | -0.912451 | 0.301951 | 0.016461 | 0.004460 | -0.006107 | 0.000491 | 0.000658 | 0.000267 | -4.775718e-04 | 0.000075 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.098990 | 2.294188 | 140° | 0.08 |
| 5 | -1.133819 | 0.771139 | 0.073925 | 0.008667 | 0.001817 | -0.000782 | -0.000424 | 0.000412 | 2.093879e-04 | 0.000050 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.073196 | 2.492090 | 140° | 0.08 |
| 6 | -0.906528 | 0.758235 | 0.025590 | 0.009584 | 0.001321 | 0.000946 | 0.001348 | 0.000506 | -1.056459e-04 | 0.000153 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.077120 | 2.287193 | 140° | 0.08 |
| 7 | -0.897786 | 0.598010 | 0.021691 | 0.007696 | -0.002021 | 0.001004 | 0.000776 | 0.000309 | -2.412954e-04 | -0.000039 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.083971 | 2.280600 | 140° | 0.08 |
| 8 | -1.069214 | 0.803718 | 0.060063 | 0.009552 | 0.002635 | -0.000024 | 0.000220 | 0.000442 | 1.188405e-04 | 0.000132 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.073029 | 2.431389 | 140° | 0.08 |
| 9 | -1.153353 | 0.201580 | 0.064673 | 0.000142 | -0.008252 | -0.002337 | -0.001004 | 0.000056 | -2.327827e-04 | -0.000266 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 2.443461 | 0.2 | 0.099243 | 2.553002 | 140° | 0.08 |
| 10 | 0.098271 | 0.584667 | -0.070353 | 0.000910 | -0.003291 | 0.000134 | -0.000159 | -0.000284 | 1.208559e-04 | -0.000146 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.089302 | 1.409695 | 80° | 0.08 |
| 11 | 0.409715 | 0.866223 | -0.078336 | -0.004074 | 0.001146 | -0.002104 | 0.000748 | 0.000136 | 2.644008e-04 | -0.000073 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.076330 | 1.242604 | 80° | 0.08 |
| 12 | 0.479149 | 1.371337 | -0.078663 | -0.005317 | 0.015571 | -0.000932 | 0.000856 | -0.001264 | -3.552821e-04 | -0.000120 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.060501 | 1.299788 | 80° | 0.08 |
| 13 | -0.043950 | 0.641566 | -0.067710 | 0.002521 | -0.002334 | 0.000573 | 0.000627 | -0.000120 | 2.669955e-05 | -0.000117 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.087227 | 1.536335 | 80° | 0.08 |
| 14 | -0.113632 | 0.414991 | -0.066985 | 0.002730 | -0.005277 | 0.000412 | 0.000743 | 0.000109 | -3.860665e-05 | 0.000190 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.098797 | 1.548685 | 80° | 0.08 |
| 15 | 0.355298 | 0.864501 | -0.078721 | -0.003226 | 0.001113 | -0.001853 | 0.000901 | 0.000137 | 2.508130e-04 | -0.000074 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.076723 | 1.280753 | 80° | 0.08 |
| 16 | -0.102059 | 0.508699 | -0.065106 | 0.002932 | -0.004082 | 0.000865 | 0.000324 | -0.000143 | -2.991154e-05 | -0.000056 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.093675 | 1.558039 | 80° | 0.08 |
| 17 | -0.121821 | 0.467478 | -0.065432 | 0.002982 | -0.004627 | 0.000701 | 0.000571 | -0.000006 | -4.302129e-05 | 0.000058 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.095911 | 1.566630 | 80° | 0.08 |
| 18 | 0.202180 | 0.844041 | -0.077693 | -0.000848 | 0.000793 | -0.001103 | 0.001261 | 0.000145 | 1.984502e-04 | -0.000068 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.078236 | 1.387295 | 80° | 0.08 |
| 19 | 0.018057 | 0.463559 | -0.069791 | 0.001618 | -0.004887 | 0.000130 | 0.000180 | -0.000058 | 8.152559e-05 | 0.000060 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 1.396263 | 0.2 | 0.095689 | 1.447270 | 80° | 0.08 |
| 20 | 1.395888 | 0.273970 | 0.029565 | -0.007117 | -0.002888 | -0.000400 | -0.000432 | 0.000188 | 2.413662e-04 | 0.000073 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.082873 | 0.390205 | 20° | 0.08 |
| 21 | 1.900065 | 0.629052 | 0.074601 | -0.016125 | 0.008819 | 0.000725 | -0.000925 | 0.000235 | -1.295535e-03 | 0.000062 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.064031 | 0.266366 | 20° | 0.08 |
| 22 | 1.312209 | 0.126744 | 0.033607 | -0.002834 | -0.004087 | 0.000589 | -0.000661 | -0.000273 | 3.846086e-04 | -0.000025 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.089630 | 0.373168 | 20° | 0.08 |
| 23 | 1.529104 | 0.313506 | 0.044290 | -0.008627 | -0.001265 | -0.000009 | -0.000039 | 0.000378 | 2.528159e-05 | 0.000031 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.078860 | 0.330521 | 20° | 0.08 |
| 24 | 1.399288 | 0.122454 | 0.045810 | -0.003021 | -0.003469 | 0.001047 | -0.000126 | -0.000142 | 2.996893e-04 | -0.000061 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.087673 | 0.318159 | 20° | 0.08 |
| 25 | 1.393050 | 0.089307 | 0.048629 | -0.002012 | -0.003558 | 0.001354 | -0.000096 | -0.000240 | 3.159606e-04 | -0.000099 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.088925 | 0.305914 | 20° | 0.08 |
| 26 | 1.510394 | 0.385975 | 0.034734 | -0.010259 | -0.000794 | -0.000713 | -0.000342 | 0.000435 | -6.106588e-07 | 0.000077 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.077160 | 0.372707 | 20° | 0.08 |
| 27 | 1.521174 | 0.167297 | 0.058388 | -0.004676 | -0.002244 | 0.001328 | 0.000450 | 0.000133 | 1.237559e-04 | -0.000085 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.083355 | 0.268017 | 20° | 0.08 |
| 28 | 1.364556 | 0.269302 | 0.025877 | -0.006852 | -0.003181 | -0.000500 | -0.000550 | 0.000141 | 2.802554e-04 | 0.000081 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.083693 | 0.406805 | 20° | 0.08 |
| 29 | 1.497570 | 0.231904 | 0.047980 | -0.006423 | -0.002178 | 0.000534 | 0.000109 | 0.000229 | 1.266806e-04 | -0.000008 | ... | 0.4 | 0.65 | 0.08 | 1.0 | 0.349066 | 0.2 | 0.081946 | 0.311726 | 20° | 0.08 |
| 30 | -0.964406 | -0.547059 | 0.002380 | -0.009928 | 0.003770 | 0.001159 | 0.001010 | 0.000236 | 1.190184e-04 | 0.000090 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.173997 | 2.326146 | 140° | 0.16 |
| 31 | -0.995482 | -0.588329 | 0.007357 | -0.011664 | 0.004883 | 0.001041 | 0.000952 | 0.000277 | 3.944896e-04 | -0.000080 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.178168 | 2.367912 | 140° | 0.16 |
| 32 | -1.084258 | -0.534348 | 0.027336 | -0.012744 | 0.001073 | -0.000244 | 0.000548 | -0.000191 | 1.501517e-04 | 0.000003 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.166143 | 2.497124 | 140° | 0.16 |
| 33 | -1.103990 | -0.332649 | 0.037067 | -0.008456 | -0.004835 | -0.001270 | 0.000250 | -0.000419 | -4.231776e-04 | 0.000217 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.142324 | 2.521001 | 140° | 0.16 |
| 34 | -1.081148 | -0.481900 | 0.029774 | -0.011638 | -0.000656 | 0.000172 | -0.000081 | -0.000507 | 1.641188e-04 | -0.000331 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.159544 | 2.492420 | 140° | 0.16 |
| 35 | -0.944329 | -0.560544 | -0.001406 | -0.009806 | 0.004606 | 0.001423 | 0.000978 | 0.000315 | 2.221716e-04 | -0.000010 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.176739 | 2.296786 | 140° | 0.16 |
| 36 | -0.947443 | -0.515159 | 0.000247 | -0.008860 | 0.002833 | 0.001258 | 0.000954 | 0.000162 | -2.426856e-05 | 0.000138 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.170455 | 2.304695 | 140° | 0.16 |
| 37 | -1.071686 | -0.381330 | 0.029611 | -0.008917 | -0.003227 | -0.000261 | 0.000127 | -0.000506 | -2.460090e-04 | -0.000004 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.148710 | 2.478740 | 140° | 0.16 |
| 38 | -1.129563 | -0.340092 | 0.042721 | -0.009278 | -0.005135 | -0.001730 | 0.000113 | -0.000464 | -3.810645e-04 | 0.000181 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.141903 | 2.555287 | 140° | 0.16 |
| 39 | -1.160450 | -0.608589 | 0.042258 | -0.017069 | 0.002021 | -0.001675 | 0.000585 | -0.000203 | 4.786634e-04 | 0.000006 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 2.443461 | 0.2 | 0.171313 | 2.610918 | 140° | 0.16 |
| 40 | -0.100311 | -0.512332 | -0.061102 | 0.000277 | 0.003444 | -0.000393 | -0.001453 | 0.000471 | 6.027916e-05 | -0.000128 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.176521 | 1.244129 | 80° | 0.16 |
| 41 | -0.079981 | -0.441534 | -0.061914 | 0.000620 | 0.001185 | -0.000248 | -0.001414 | 0.000194 | -1.683988e-05 | 0.000093 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.167533 | 1.252522 | 80° | 0.16 |
| 42 | -0.259471 | -0.477805 | -0.061752 | -0.000556 | 0.003429 | 0.000534 | -0.000928 | 0.000441 | -6.780287e-06 | -0.000085 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.175097 | 1.438110 | 80° | 0.16 |
| 43 | -0.302468 | -0.225359 | -0.060633 | 0.001001 | -0.002737 | 0.001199 | -0.000374 | -0.000215 | -2.950729e-04 | 0.000315 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.146855 | 1.570428 | 80° | 0.16 |
| 44 | -0.152787 | -0.194943 | -0.065320 | 0.001167 | -0.003807 | 0.000371 | -0.000481 | -0.000161 | -2.123430e-04 | 0.000508 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.143005 | 1.422087 | 80° | 0.16 |
| 45 | -0.307667 | -0.469121 | -0.060844 | -0.000789 | 0.003396 | 0.000805 | -0.000743 | 0.000429 | -3.411932e-05 | -0.000063 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.174511 | 1.497144 | 80° | 0.16 |
| 46 | -0.275157 | -0.439585 | -0.061692 | -0.000384 | 0.002311 | 0.000701 | -0.000792 | 0.000313 | -8.908570e-05 | 0.000045 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.170496 | 1.470243 | 80° | 0.16 |
| 47 | -0.052624 | -0.296443 | -0.062555 | 0.001237 | -0.002283 | 0.000227 | -0.001640 | -0.000360 | -4.048301e-07 | 0.000094 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.151212 | 1.282264 | 80° | 0.16 |
| 48 | -0.112605 | -0.512113 | -0.061268 | 0.000202 | 0.003539 | -0.000323 | -0.001436 | 0.000476 | 6.406333e-05 | -0.000145 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.176779 | 1.257857 | 80° | 0.16 |
| 49 | -0.274825 | -0.335715 | -0.059410 | 0.000581 | -0.000319 | 0.001512 | -0.001492 | -0.000330 | -3.504779e-05 | -0.000270 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 1.396263 | 0.2 | 0.157964 | 1.505475 | 80° | 0.16 |
| 50 | 0.666826 | -0.751041 | 0.016709 | 0.010780 | 0.002460 | -0.000799 | 0.000803 | 0.000095 | -1.436529e-04 | 0.000089 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.166466 | 0.346456 | 20° | 0.16 |
| 51 | 0.578040 | -0.806088 | 0.008922 | 0.010286 | 0.005021 | -0.001514 | 0.000503 | 0.000469 | -2.606051e-04 | -0.000155 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.177635 | 0.386719 | 20° | 0.16 |
| 52 | 0.653467 | -0.624203 | 0.006614 | 0.009087 | -0.000103 | -0.000554 | -0.000854 | -0.000533 | 1.463360e-04 | -0.000157 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.155776 | 0.441191 | 20° | 0.16 |
| 53 | 0.572528 | -0.783451 | 0.005996 | 0.009685 | 0.004365 | -0.001549 | 0.000388 | 0.000436 | -2.251687e-04 | -0.000036 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.175787 | 0.407493 | 20° | 0.16 |
| 54 | 0.659641 | -0.513237 | -0.004183 | 0.006170 | -0.002880 | -0.000997 | -0.000403 | -0.000293 | 2.174160e-04 | 0.000516 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.146865 | 0.504861 | 20° | 0.16 |
| 55 | 0.608625 | -0.678892 | 0.002987 | 0.008683 | 0.001219 | -0.001112 | -0.000207 | -0.000031 | -6.913243e-06 | 0.000121 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.163440 | 0.445301 | 20° | 0.16 |
| 56 | 0.686187 | -0.696469 | 0.015975 | 0.010500 | 0.001120 | -0.000493 | 0.000174 | -0.000254 | 3.747075e-06 | -0.000045 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.160243 | 0.366476 | 20° | 0.16 |
| 57 | 0.706113 | -0.689142 | 0.018765 | 0.010884 | 0.000886 | -0.000274 | 0.000101 | -0.000391 | 3.554720e-05 | -0.000137 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.158335 | 0.354346 | 20° | 0.16 |
| 58 | 0.863564 | -0.675819 | 0.039638 | 0.012348 | -0.000697 | 0.000855 | 0.001824 | -0.000443 | 5.866481e-05 | 0.000142 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.148304 | 0.235357 | 20° | 0.16 |
| 59 | 0.804667 | -0.784796 | 0.042667 | 0.014671 | 0.002395 | 0.000714 | 0.001847 | -0.000351 | -1.425091e-04 | -0.000275 | ... | 0.4 | 0.65 | 0.16 | 1.0 | 0.349066 | 0.2 | 0.160054 | 0.206858 | 20° | 0.16 |
60 rows × 22 columns
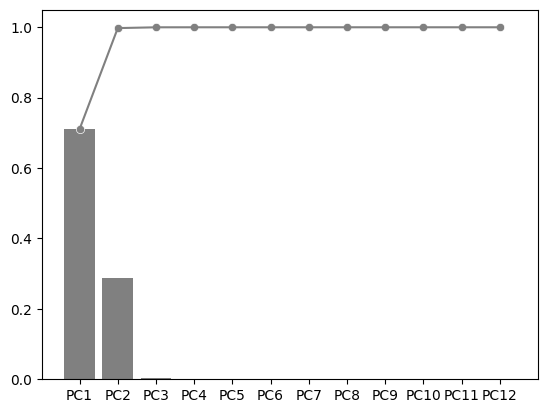
fig, ax = plt.subplots()
explained_variance_ratio_plot(pca, ax=ax, verbose=True)
PC1 PC2 PC3 PC4 PC5 PC6 PC7 PC8 PC9 PC10 PC11 PC12
Explained Variance Ratio (EVR) 0.7105 0.2872 0.0023 0.0001 0.0000 0.0000 0.0000 0.0000 0.0000 0.0000 0.0000 0.0000
Cumsum of EVR 0.7105 0.9977 0.9999 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000
<Axes: >
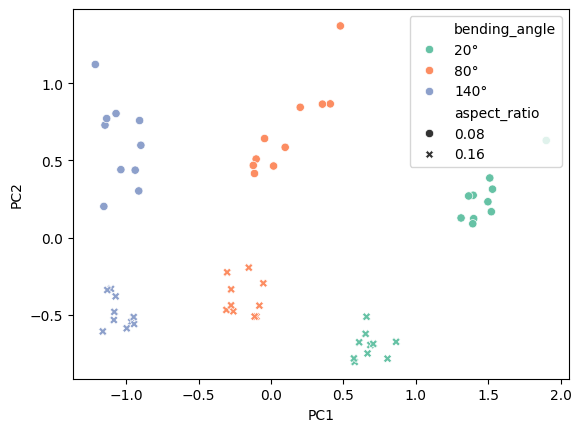
fig, ax = plt.subplots()
sns.scatterplot(
data=df_pca,
x="PC1",
y="PC2",
hue="bending_angle",
hue_order=bending_order,
style="aspect_ratio",
palette="Set2",
ax=ax,
)
<Axes: xlabel='PC1', ylabel='PC2'>
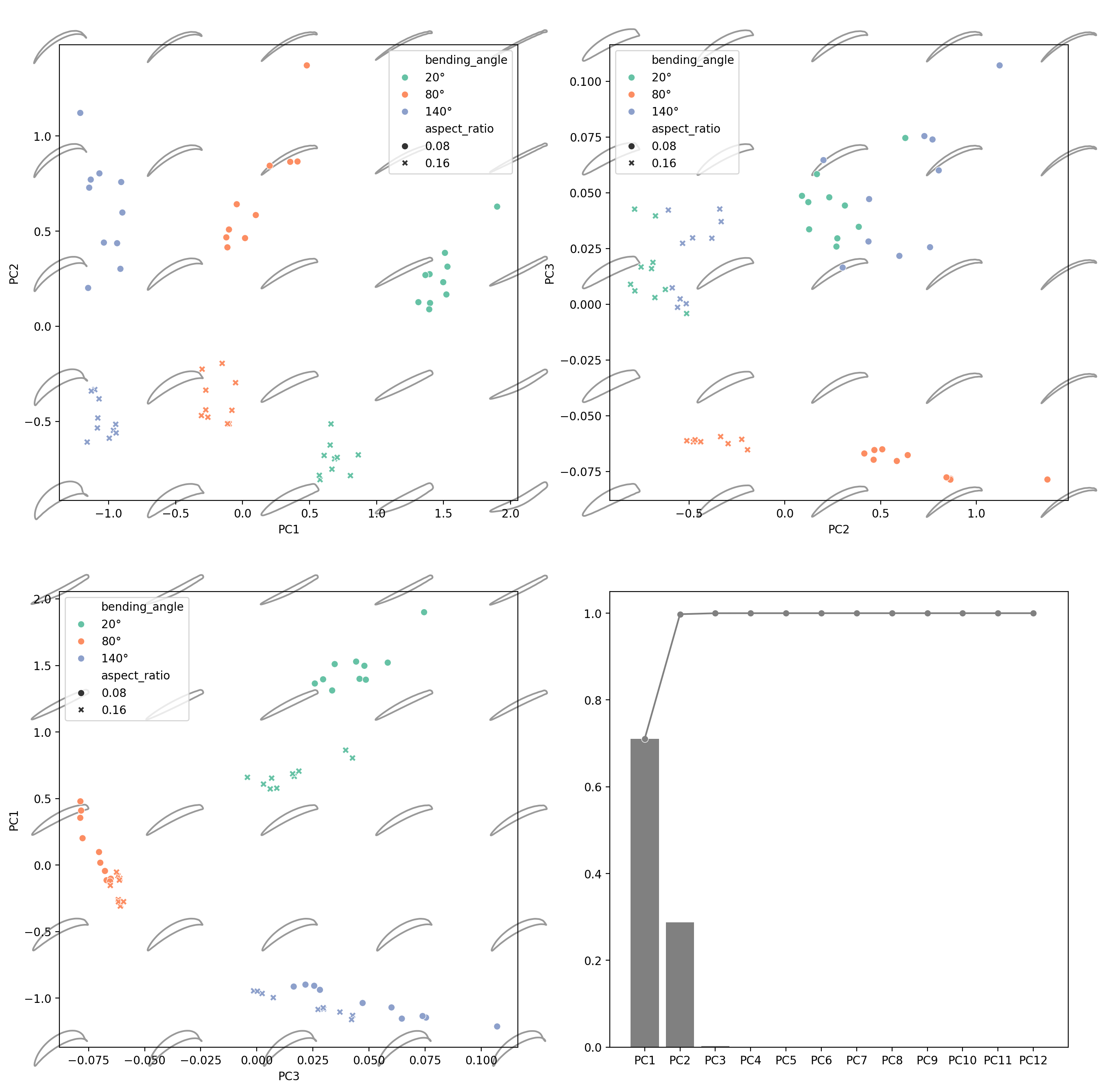
Morphospace#
def render_curve_3d_fixed_view(coords, ax, *, color="gray", alpha=0.7, **kw):
kw.pop("links", None)
ax.plot(coords[:, 0], coords[:, 1], coords[:, 2], color=color, alpha=alpha)
ptp = [max(np.max(coords[:, i]) - np.min(coords[:, i]), 1e-10) for i in range(3)]
ax.set_box_aspect(ptp)
ax.view_init(elev=60, azim=-120)
fig, axes = plt.subplots(2, 2, figsize=(16, 16), dpi=200)
for ax, (i, j) in zip(axes.flat[:3], [(0, 1), (1, 2), (2, 0)]):
morphospace_plot(
data=df_pca,
x=f"PC{i + 1}", y=f"PC{j + 1}",
hue="bending_angle", hue_order=bending_order,
palette="Set2",
reducer=pca,
descriptor=efa,
shape_type="curve_3d",
render_fn=render_curve_3d_fixed_view,
components=(i, j),
n_shapes=5,
shape_scale=0.8,
shape_color="gray",
shape_alpha=0.8,
ax=ax,
scatter_kw=dict(style="aspect_ratio"),
)
explained_variance_ratio_plot(pca, ax=axes[1, 1], verbose=True)
PC1 PC2 PC3 PC4 PC5 PC6 PC7 PC8 PC9 PC10 PC11 PC12
Explained Variance Ratio (EVR) 0.7105 0.2872 0.0023 0.0001 0.0000 0.0000 0.0000 0.0000 0.0000 0.0000 0.0000 0.0000
Cumsum of EVR 0.7105 0.9977 0.9999 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000
<Axes: >