Elliptic Fourier Analysis#
See also
For cases where automatic normalization is not suitable, see 2D Outline Registration.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.decomposition import PCA
from ktch.datasets import load_outline_mosquito_wings
from ktch.harmonic import EllipticFourierAnalysis
from ktch.plot import explained_variance_ratio_plot, morphospace_plot, shape_variation_plot
Load mosquito wing outline dataset#
from Rohlf and Archie 1984 Syst. Zool.
data_outline_mosquito_wings = load_outline_mosquito_wings(as_frame=True)
data_outline_mosquito_wings.coords
| x | y | ||
|---|---|---|---|
| specimen_id | coord_id | ||
| 0 | 0 | 0.99973 | 0.00000 |
| 1 | 0.81881 | 0.05151 | |
| 2 | 0.70063 | 0.08851 | |
| 3 | 0.61179 | 0.11670 | |
| 4 | 0.53226 | 0.13666 | |
| ... | ... | ... | ... |
| 125 | 95 | 0.68167 | -0.22149 |
| 96 | 0.76135 | -0.19548 | |
| 97 | 0.90752 | -0.17312 | |
| 98 | 0.95720 | -0.12093 | |
| 99 | 0.95964 | -0.06038 |
12600 rows × 2 columns
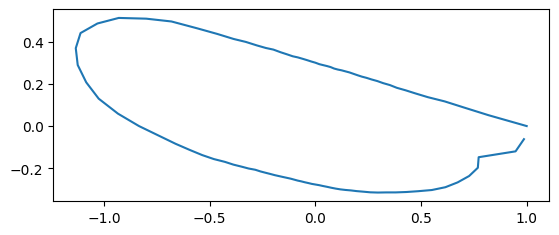
coords = data_outline_mosquito_wings.coords.to_numpy().reshape(-1, 100, 2)
fig, ax = plt.subplots()
sns.lineplot(x=coords[0][:, 0], y=coords[0][:, 1], sort=False, estimator=None, ax=ax)
ax.set_aspect("equal")
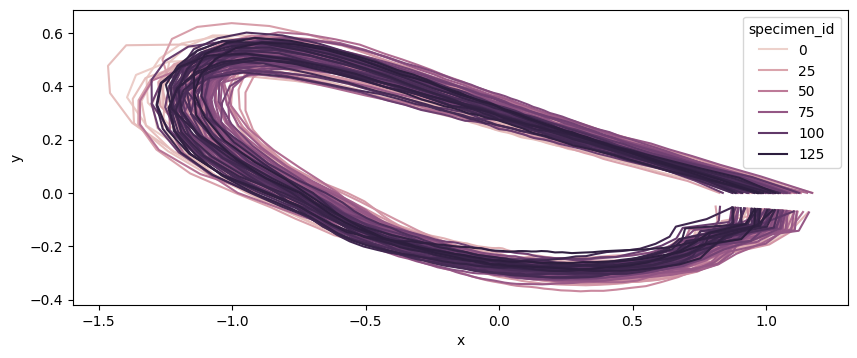
fig, ax = plt.subplots(figsize=(10, 10))
sns.lineplot(
data=data_outline_mosquito_wings.coords,
x="x",
y="y",
hue="specimen_id",
sort=False,
estimator=None,
ax=ax,
)
ax.set_aspect("equal")
EFA#
efa = EllipticFourierAnalysis(n_harmonics=20)
coef = efa.fit_transform(coords)
PCA#
pca = PCA(n_components=12)
pcscores = pca.fit_transform(coef)
df_pca = pd.DataFrame(pcscores)
df_pca["specimen_id"] = list(range(len(pcscores)))
df_pca = df_pca.set_index("specimen_id")
df_pca = df_pca.join(data_outline_mosquito_wings.meta)
df_pca = df_pca.rename(columns={i: ("PC" + str(i + 1)) for i in range(12)})
df_pca
| PC1 | PC2 | PC3 | PC4 | PC5 | PC6 | PC7 | PC8 | PC9 | PC10 | PC11 | PC12 | genus | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| specimen_id | |||||||||||||
| 0 | 0.004277 | -0.019211 | 0.018697 | 0.000810 | -0.001363 | -0.015705 | 0.007489 | -0.000853 | -0.004332 | 0.000064 | -0.001233 | -0.000509 | AN |
| 1 | -0.034207 | 0.019829 | 0.000914 | 0.002444 | 0.004912 | 0.004622 | 0.002922 | -0.000719 | -0.008601 | -0.003872 | -0.002954 | 0.000656 | AN |
| 2 | -0.199464 | -0.031161 | -0.045073 | 0.008504 | -0.004950 | 0.006258 | 0.001273 | 0.004424 | -0.003027 | -0.000986 | 0.002781 | -0.001875 | AN |
| 3 | -0.248697 | 0.035792 | -0.019214 | -0.000370 | 0.007445 | -0.009597 | 0.004925 | 0.001945 | 0.001729 | -0.001801 | 0.001553 | -0.001683 | AN |
| 4 | 0.151898 | -0.057919 | -0.009257 | 0.009022 | 0.002941 | 0.009306 | 0.003443 | -0.001652 | -0.001026 | -0.004647 | -0.003179 | 0.000502 | AN |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 121 | -0.084827 | 0.032089 | -0.023891 | 0.000607 | 0.007980 | -0.003300 | 0.000688 | 0.002950 | 0.003434 | -0.001497 | 0.002059 | 0.000181 | CX |
| 122 | -0.096891 | -0.043715 | 0.000709 | -0.007439 | -0.005802 | -0.003579 | 0.005781 | 0.000594 | 0.006692 | 0.001221 | -0.000017 | 0.001236 | CX |
| 123 | -0.119387 | 0.016518 | -0.031522 | -0.021842 | 0.001695 | 0.002130 | 0.002902 | 0.003965 | 0.000240 | -0.001915 | -0.001795 | 0.000932 | CX |
| 124 | -0.059915 | 0.046869 | 0.014093 | -0.002840 | 0.008363 | -0.001264 | 0.001011 | 0.000412 | 0.003208 | 0.004421 | -0.000136 | 0.000643 | CX |
| 125 | -0.035477 | 0.031945 | 0.008483 | -0.019398 | -0.002612 | 0.005169 | 0.004838 | -0.000205 | 0.002391 | -0.002302 | 0.000625 | -0.001556 | DE |
126 rows × 13 columns
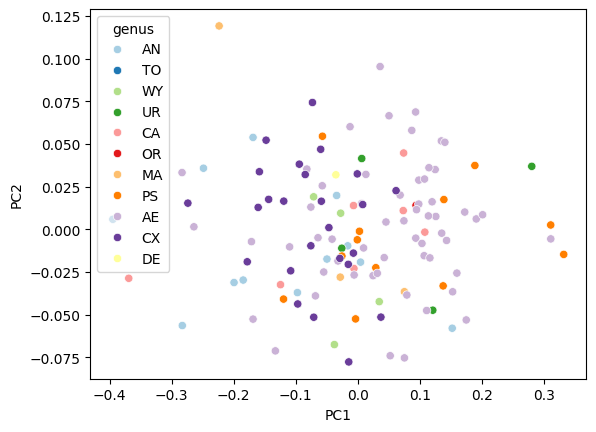
fig, ax = plt.subplots()
sns.scatterplot(data=df_pca, x="PC1", y="PC2", hue="genus", ax=ax, palette="Paired")
<Axes: xlabel='PC1', ylabel='PC2'>
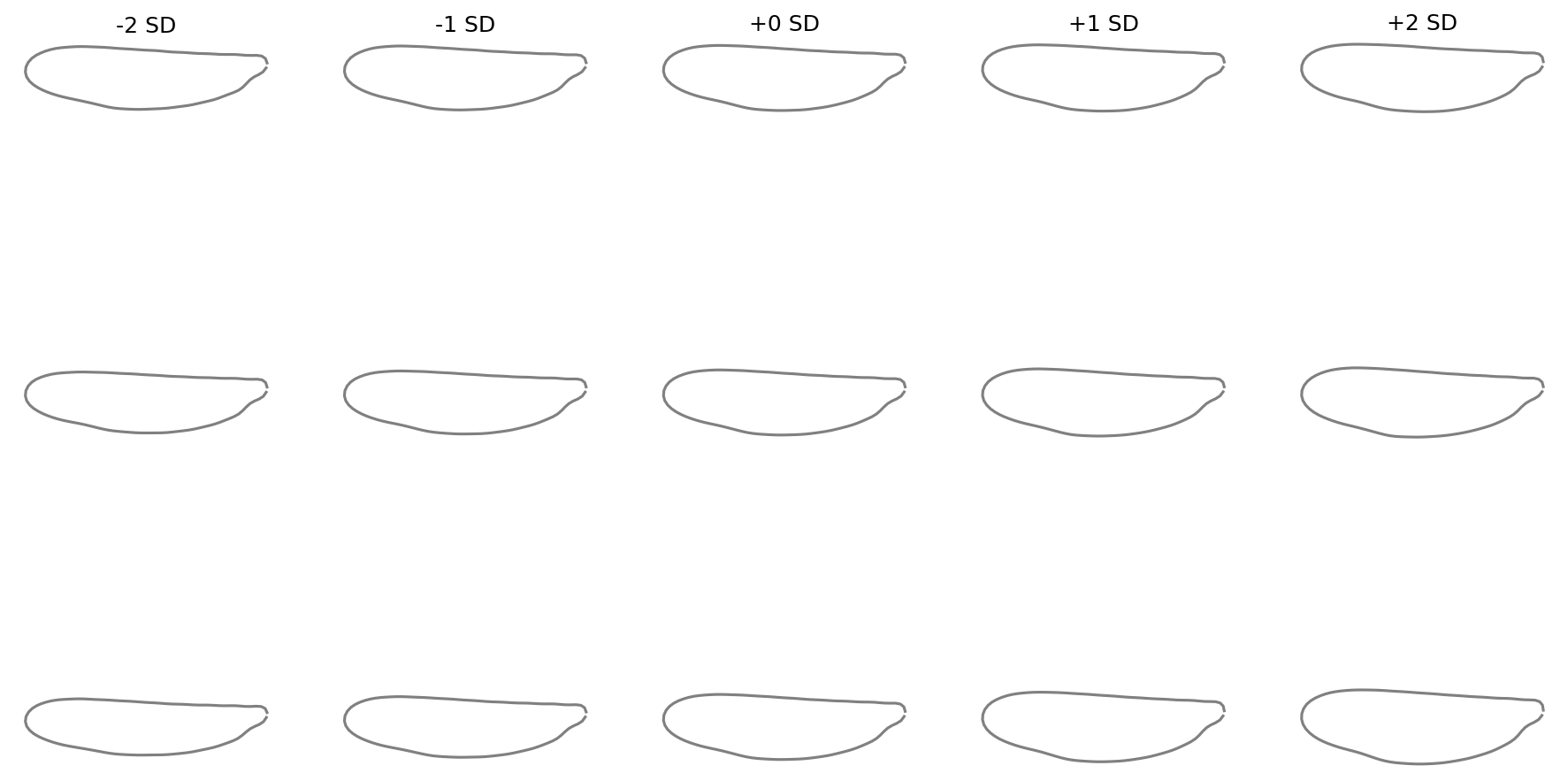
Shape variation along PC axes#
fig = shape_variation_plot(
pca,
descriptor=efa,
components=(0, 1, 2),
sd_values=(-2, -1, 0, 1, 2),
)
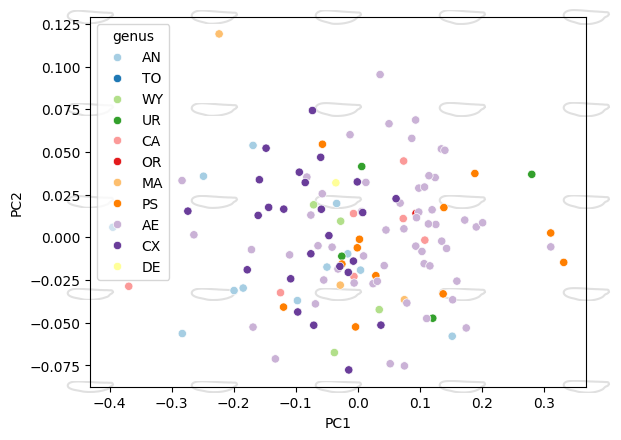
Morphospace#
ax = morphospace_plot(
data=df_pca,
x="PC1", y="PC2", hue="genus",
reducer=pca,
descriptor=efa,
palette="Paired",
n_shapes=5,
shape_scale=0.5,
)
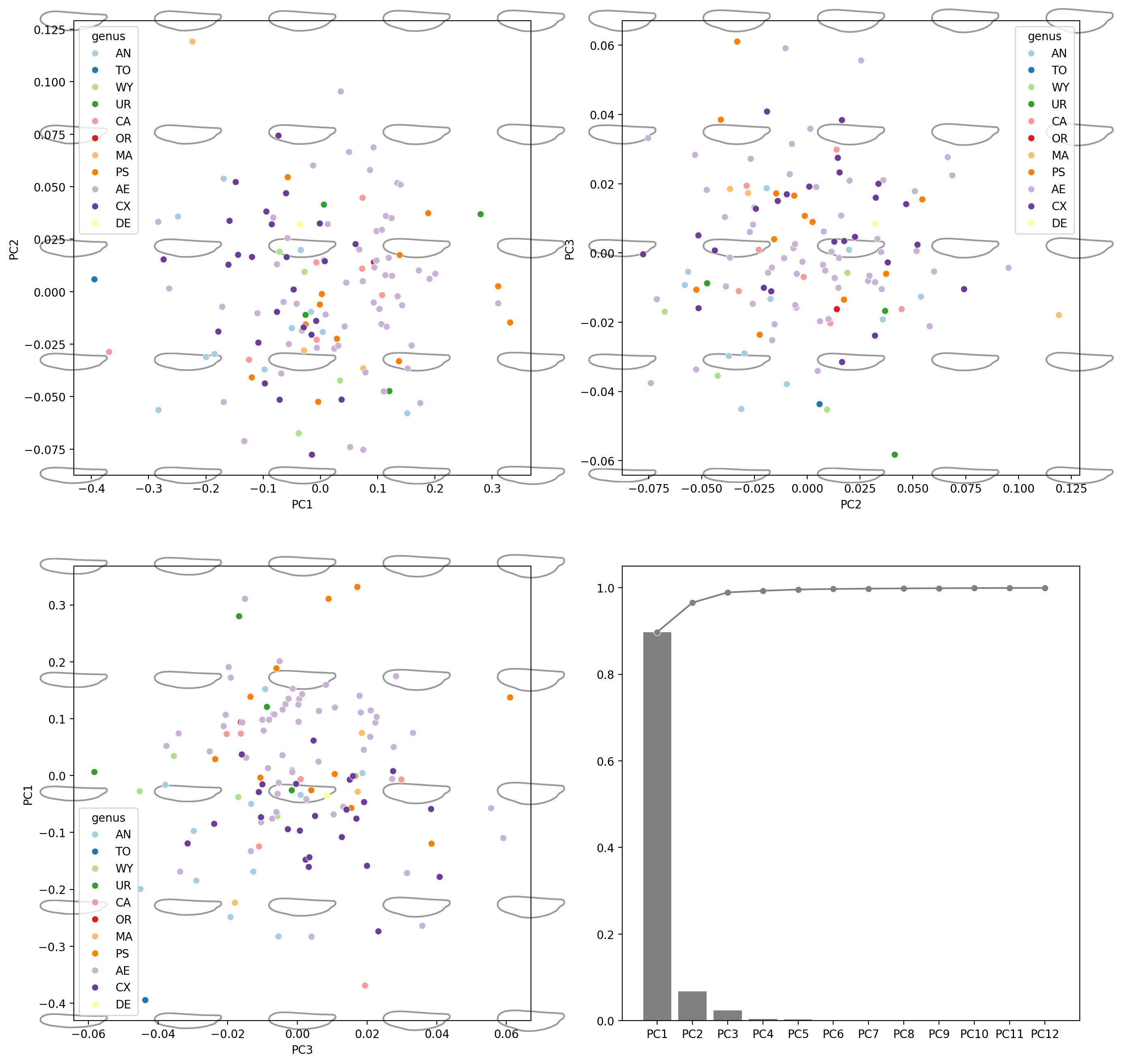
fig, axes = plt.subplots(2, 2, figsize=(16, 16), dpi=200)
for ax, (i, j) in zip(axes.flat[:3], [(0, 1), (1, 2), (2, 0)]):
morphospace_plot(
data=df_pca,
x=f"PC{i + 1}", y=f"PC{j + 1}", hue="genus",
reducer=pca,
descriptor=efa,
components=(i, j),
palette="Paired",
n_shapes=5,
shape_color="gray",
shape_scale=0.8,
shape_alpha=0.8,
ax=ax,
)
explained_variance_ratio_plot(pca, ax=axes[1, 1])
<Axes: >