Generalized Procrustes analysis#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.collections import LineCollection
import seaborn as sns
from sklearn.decomposition import PCA
from ktch.datasets import load_landmark_mosquito_wings
from ktch.landmark import GeneralizedProcrustesAnalysis
from ktch.plot import explained_variance_ratio_plot, morphospace_plot, shape_variation_plot
Load mosquito wing landmark dataset#
from Rohlf and Slice 1990 Syst. Zool.
data_landmark_mosquito_wings = load_landmark_mosquito_wings(as_frame=True)
data_landmark_mosquito_wings.coords
| x | y | ||
|---|---|---|---|
| specimen_id | coord_id | ||
| 0 | 0 | -0.4933 | 0.0130 |
| 1 | -0.0777 | 0.0832 | |
| 2 | 0.2231 | 0.0861 | |
| 3 | 0.2641 | 0.0462 | |
| 4 | 0.2645 | 0.0261 | |
| ... | ... | ... | ... |
| 126 | 13 | -0.2028 | 0.0371 |
| 14 | 0.0490 | 0.0347 | |
| 15 | -0.0422 | 0.0204 | |
| 16 | 0.1004 | -0.0180 | |
| 17 | -0.1473 | -0.0057 |
2286 rows × 2 columns
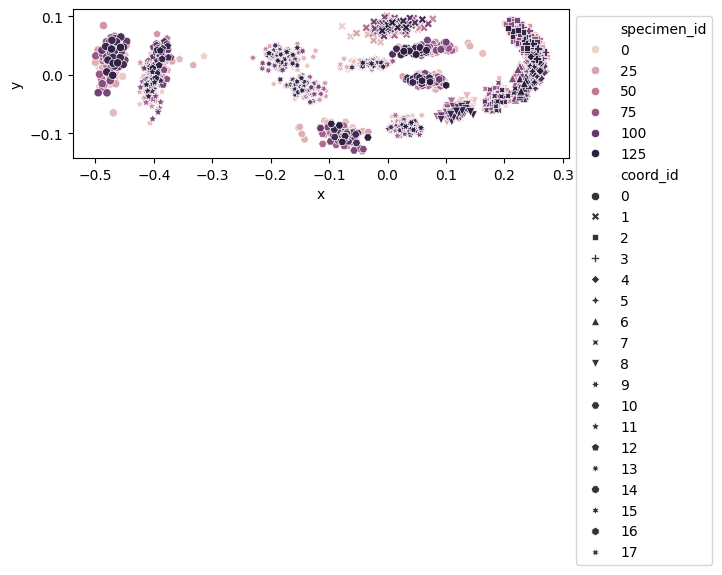
fig, ax = plt.subplots()
sns.scatterplot(
data=data_landmark_mosquito_wings.coords,
x="x",
y="y",
hue="specimen_id",
style="coord_id",
ax=ax,
)
ax.set_aspect("equal")
ax.legend(loc="upper left", bbox_to_anchor=(1, 1))
<matplotlib.legend.Legend at 0x7fbbf4b4d350>
For applying generalized Procrustes analysis (GPA), we convert the configuration data into DataFrame of shape n_specimens x (n_landmarks*n_dim).
def configuration_plot(
configuration_2d,
x="x",
y="y",
links=None,
ax=None,
hue=None,
hue_order=None,
c="gray",
palette=None,
c_line="gray",
style=None,
s=10,
alpha=1,
):
if ax is None:
fig = plt.figure()
ax = fig.add_subplot(111)
if links is None:
links = []
configuration = configuration_2d.reset_index()
if hue is not None and hue_order is None:
hue_order = configuration[hue].unique()
color_map = None
if hue is not None:
if palette is not None:
colors = sns.color_palette(palette, n_colors=len(hue_order))
color_map = dict(zip(hue_order, colors))
else:
hue_dtype = configuration[hue].dtype
if np.issubdtype(hue_dtype, np.number):
cmap = sns.cubehelix_palette(as_cmap=True)
hue_min, hue_max = min(hue_order), max(hue_order)
color_map = {}
for hue_val in hue_order:
if hue_max > hue_min:
norm_val = (hue_val - hue_min) / (hue_max - hue_min)
else:
norm_val = 0.5
color_map[hue_val] = cmap(norm_val)
else:
colors = sns.color_palette(n_colors=len(hue_order))
color_map = dict(zip(hue_order, colors))
if links:
if hue is None:
segments = []
for link in links:
link_data = configuration[configuration["coord_id"].isin(link)]
if len(link_data) == 2:
coords = link_data[[x, y]].values
segments.append(coords)
if segments:
lc = LineCollection(segments, colors=c_line, alpha=alpha)
ax.add_collection(lc)
else:
for specimen in hue_order:
specimen_data = configuration[configuration[hue] == specimen]
segments = []
for link in links:
link_data = specimen_data[specimen_data["coord_id"].isin(link)]
if len(link_data) == 2:
coords = link_data[[x, y]].values
segments.append(coords)
if segments:
lc = LineCollection(
segments, colors=[color_map[specimen]], alpha=alpha
)
ax.add_collection(lc)
ax.autoscale_view()
axis = sns.scatterplot(
data=configuration,
x=x,
y=y,
ax=ax,
hue=hue,
hue_order=hue_order,
palette=palette,
style=style,
c=c,
alpha=alpha,
s=s,
)
if axis.legend_:
sns.move_legend(ax, "upper left", bbox_to_anchor=(1, 1))
ax.set_aspect("equal")
links = [
[0, 1],
[1, 2],
[2, 3],
[3, 4],
[4, 5],
[5, 6],
[6, 7],
[7, 8],
[8, 9],
[9, 10],
[10, 11],
[1, 12],
[2, 12],
[13, 14],
[14, 3],
[14, 4],
[15, 5],
[16, 6],
[16, 7],
[17, 8],
[17, 9],
]
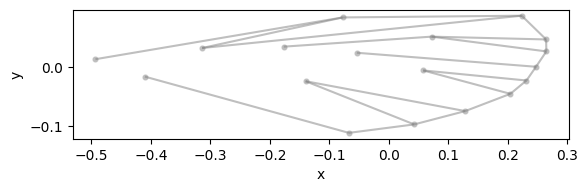
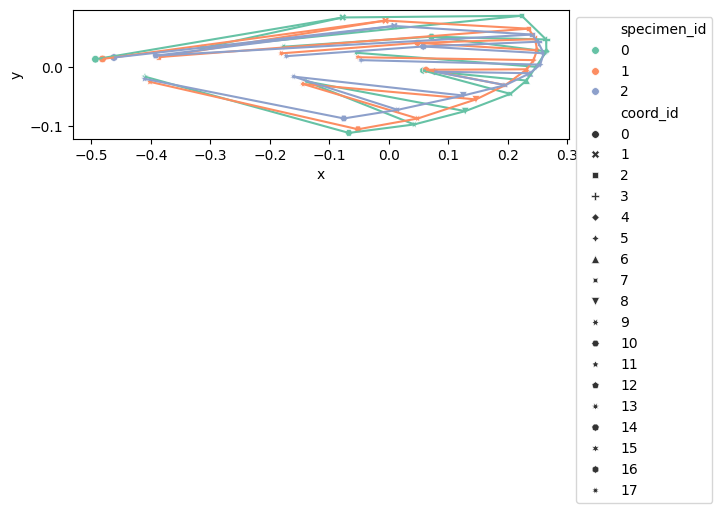
configuration_plot(
data_landmark_mosquito_wings.coords.loc[0], links=links, alpha=0.5, s=20
)
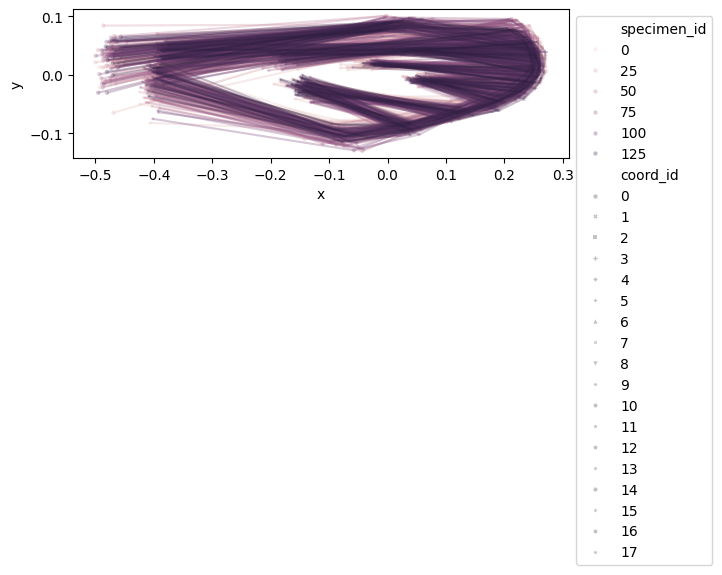
configuration_plot(
data_landmark_mosquito_wings.coords,
links=links,
alpha=0.3,
hue="specimen_id",
style="coord_id",
)
configuration_plot(
data_landmark_mosquito_wings.coords.loc[0:2],
links=links,
hue="specimen_id",
style="coord_id",
palette="Set2",
s=30,
)
index = pd.MultiIndex.from_tuples(
[(1, i) for i in data_landmark_mosquito_wings.coords.loc[1].index],
names=["specimen_id", "coord_id"],
)
x2 = pd.DataFrame(
data_landmark_mosquito_wings.coords.loc[1].to_numpy(),
columns=["x", "y"],
index=index,
)
data_landmark_mosquito_wings.coords.loc[0:2]
| x | y | ||
|---|---|---|---|
| specimen_id | coord_id | ||
| 0 | 0 | -0.4933 | 0.0130 |
| 1 | -0.0777 | 0.0832 | |
| 2 | 0.2231 | 0.0861 | |
| 3 | 0.2641 | 0.0462 | |
| 4 | 0.2645 | 0.0261 | |
| 5 | 0.2471 | 0.0003 | |
| 6 | 0.2311 | -0.0228 | |
| 7 | 0.2040 | -0.0452 | |
| 8 | 0.1282 | -0.0742 | |
| 9 | 0.0424 | -0.0966 | |
| 10 | -0.0674 | -0.1108 | |
| 11 | -0.4102 | -0.0163 | |
| 12 | -0.3140 | 0.0318 | |
| 13 | -0.1768 | 0.0341 | |
| 14 | 0.0715 | 0.0509 | |
| 15 | -0.0540 | 0.0238 | |
| 16 | 0.0575 | -0.0059 | |
| 17 | -0.1401 | -0.0240 | |
| 1 | 0 | -0.4814 | 0.0135 |
| 1 | -0.0058 | 0.0780 | |
| 2 | 0.2345 | 0.0644 | |
| 3 | 0.2460 | 0.0467 | |
| 4 | 0.2487 | 0.0281 | |
| 5 | 0.2430 | 0.0115 | |
| 6 | 0.2316 | -0.0039 | |
| 7 | 0.1956 | -0.0305 | |
| 8 | 0.1462 | -0.0545 | |
| 9 | 0.0483 | -0.0866 | |
| 10 | -0.0520 | -0.1047 | |
| 11 | -0.4016 | -0.0250 | |
| 12 | -0.3868 | 0.0166 | |
| 13 | -0.1808 | 0.0229 | |
| 14 | 0.0484 | 0.0405 | |
| 15 | -0.0519 | 0.0164 | |
| 16 | 0.0623 | -0.0047 | |
| 17 | -0.1444 | -0.0286 | |
| 2 | 0 | -0.4622 | 0.0159 |
| 1 | 0.0089 | 0.0689 | |
| 2 | 0.2404 | 0.0545 | |
| 3 | 0.2501 | 0.0424 | |
| 4 | 0.2600 | 0.0230 | |
| 5 | 0.2541 | 0.0039 | |
| 6 | 0.2369 | -0.0105 | |
| 7 | 0.1957 | -0.0305 | |
| 8 | 0.1249 | -0.0480 | |
| 9 | 0.0146 | -0.0720 | |
| 10 | -0.0758 | -0.0865 | |
| 11 | -0.4104 | -0.0200 | |
| 12 | -0.3919 | 0.0190 | |
| 13 | -0.1724 | 0.0182 | |
| 14 | 0.0577 | 0.0344 | |
| 15 | -0.0468 | 0.0115 | |
| 16 | 0.0766 | -0.0079 | |
| 17 | -0.1602 | -0.0162 |
df_coords = (
data_landmark_mosquito_wings.coords.unstack()
.swaplevel(1, 0, axis=1)
.sort_index(axis=1)
)
df_coords.columns = [
dim + "_" + str(landmark_idx) for landmark_idx, dim in df_coords.columns
]
df_coords
| x_0 | y_0 | x_1 | y_1 | x_2 | y_2 | x_3 | y_3 | x_4 | y_4 | ... | x_13 | y_13 | x_14 | y_14 | x_15 | y_15 | x_16 | y_16 | x_17 | y_17 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| specimen_id | |||||||||||||||||||||
| 0 | -0.4933 | 0.0130 | -0.0777 | 0.0832 | 0.2231 | 0.0861 | 0.2641 | 0.0462 | 0.2645 | 0.0261 | ... | -0.1768 | 0.0341 | 0.0715 | 0.0509 | -0.0540 | 0.0238 | 0.0575 | -0.0059 | -0.1401 | -0.0240 |
| 1 | -0.4814 | 0.0135 | -0.0058 | 0.0780 | 0.2345 | 0.0644 | 0.2460 | 0.0467 | 0.2487 | 0.0281 | ... | -0.1808 | 0.0229 | 0.0484 | 0.0405 | -0.0519 | 0.0164 | 0.0623 | -0.0047 | -0.1444 | -0.0286 |
| 2 | -0.4622 | 0.0159 | 0.0089 | 0.0689 | 0.2404 | 0.0545 | 0.2501 | 0.0424 | 0.2600 | 0.0230 | ... | -0.1724 | 0.0182 | 0.0577 | 0.0344 | -0.0468 | 0.0115 | 0.0766 | -0.0079 | -0.1602 | -0.0162 |
| 3 | -0.4534 | -0.0028 | -0.0318 | 0.0738 | 0.2423 | 0.0808 | 0.2627 | 0.0559 | 0.2654 | 0.0322 | ... | -0.1536 | 0.0150 | 0.0617 | 0.0436 | -0.0549 | 0.0217 | 0.0705 | -0.0031 | -0.1507 | -0.0273 |
| 4 | -0.4926 | -0.0212 | -0.0260 | 0.0708 | 0.2347 | 0.0679 | 0.2398 | 0.0584 | 0.2415 | 0.0355 | ... | -0.1374 | 0.0154 | 0.0762 | 0.0457 | -0.0313 | 0.0170 | 0.0880 | 0.0037 | -0.1318 | -0.0278 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 122 | -0.4703 | -0.0009 | 0.0011 | 0.0805 | 0.2146 | 0.0747 | 0.2348 | 0.0585 | 0.2453 | 0.0350 | ... | -0.1624 | 0.0193 | 0.0557 | 0.0467 | -0.0272 | 0.0190 | 0.0718 | -0.0052 | -0.1583 | -0.0251 |
| 123 | -0.4725 | 0.0441 | 0.0318 | 0.0808 | 0.2382 | 0.0419 | 0.2504 | 0.0237 | 0.2492 | 0.0019 | ... | -0.1724 | 0.0390 | 0.0667 | 0.0418 | -0.0108 | 0.0205 | 0.0548 | -0.0049 | -0.1546 | -0.0115 |
| 124 | -0.4697 | 0.0196 | 0.0007 | 0.0850 | 0.2228 | 0.0760 | 0.2485 | 0.0540 | 0.2548 | 0.0319 | ... | -0.1816 | 0.0213 | 0.0376 | 0.0408 | -0.0073 | 0.0137 | 0.0592 | -0.0103 | -0.1531 | -0.0300 |
| 125 | -0.4620 | 0.0204 | 0.0082 | 0.0893 | 0.2061 | 0.0797 | 0.2374 | 0.0529 | 0.2457 | 0.0306 | ... | -0.1876 | 0.0241 | 0.0279 | 0.0403 | -0.0425 | 0.0145 | 0.0729 | -0.0123 | -0.1460 | -0.0269 |
| 126 | -0.4570 | 0.0468 | -0.0090 | 0.0753 | 0.2381 | 0.0463 | 0.2556 | 0.0275 | 0.2614 | 0.0064 | ... | -0.2028 | 0.0371 | 0.0490 | 0.0347 | -0.0422 | 0.0204 | 0.1004 | -0.0180 | -0.1473 | -0.0057 |
127 rows × 36 columns
GPA#
gpa = GeneralizedProcrustesAnalysis().set_output(transform="pandas")
df_shapes = gpa.fit_transform(df_coords)
df_shapes
| x_0 | y_0 | x_1 | y_1 | x_2 | y_2 | x_3 | y_3 | x_4 | y_4 | ... | x_13 | y_13 | x_14 | y_14 | x_15 | y_15 | x_16 | y_16 | x_17 | y_17 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| specimen_id | |||||||||||||||||||||
| 0 | -0.488913 | 0.021159 | -0.075651 | 0.083813 | 0.222655 | 0.081656 | 0.262641 | 0.041408 | 0.262701 | 0.021471 | ... | -0.174735 | 0.036786 | 0.071747 | 0.049290 | -0.053145 | 0.024519 | 0.056916 | -0.006796 | -0.139317 | -0.021437 |
| 1 | -0.479850 | 0.030667 | -0.002995 | 0.078029 | 0.236289 | 0.055873 | 0.247131 | 0.037801 | 0.249161 | 0.019145 | ... | -0.179578 | 0.029304 | 0.049746 | 0.038675 | -0.051194 | 0.018213 | 0.062000 | -0.006922 | -0.145098 | -0.023383 |
| 2 | -0.461238 | 0.021194 | 0.009668 | 0.068674 | 0.240606 | 0.051631 | 0.250150 | 0.039440 | 0.259809 | 0.019959 | ... | -0.171907 | 0.020150 | 0.057987 | 0.033671 | -0.046599 | 0.012014 | 0.076367 | -0.008775 | -0.160124 | -0.014331 |
| 3 | -0.451582 | 0.021733 | -0.027671 | 0.075199 | 0.245617 | 0.067345 | 0.264582 | 0.041450 | 0.265988 | 0.017707 | ... | -0.152122 | 0.023242 | 0.063790 | 0.040074 | -0.053488 | 0.024575 | 0.070026 | -0.006899 | -0.151522 | -0.019031 |
| 4 | -0.491092 | 0.019428 | -0.020015 | 0.072475 | 0.238724 | 0.048169 | 0.243009 | 0.038313 | 0.242816 | 0.015425 | ... | -0.135231 | 0.026596 | 0.079447 | 0.039142 | -0.029701 | 0.019466 | 0.087717 | -0.003550 | -0.133219 | -0.016779 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 122 | -0.468838 | 0.028646 | 0.006143 | 0.080164 | 0.218584 | 0.060969 | 0.237700 | 0.043553 | 0.246690 | 0.019469 | ... | -0.160671 | 0.029436 | 0.058442 | 0.043044 | -0.025929 | 0.020642 | 0.071228 | -0.009699 | -0.159374 | -0.015077 |
| 123 | -0.473804 | 0.016670 | 0.027045 | 0.082425 | 0.235150 | 0.055559 | 0.248370 | 0.038112 | 0.248433 | 0.016300 | ... | -0.174200 | 0.028931 | 0.064108 | 0.045546 | -0.011957 | 0.019822 | 0.054939 | -0.001719 | -0.153528 | -0.020407 |
| 124 | -0.468449 | 0.032047 | 0.002958 | 0.084844 | 0.224473 | 0.069955 | 0.249549 | 0.047306 | 0.255251 | 0.025073 | ... | -0.180752 | 0.026088 | 0.038626 | 0.039732 | -0.006925 | 0.013867 | 0.058834 | -0.011863 | -0.153659 | -0.025890 |
| 125 | -0.459471 | 0.036434 | 0.011277 | 0.088662 | 0.208061 | 0.072197 | 0.238303 | 0.044411 | 0.245792 | 0.021909 | ... | -0.186025 | 0.030549 | 0.029190 | 0.039168 | -0.041832 | 0.015925 | 0.072178 | -0.014794 | -0.146368 | -0.021701 |
| 126 | -0.457745 | 0.025260 | -0.012492 | 0.074647 | 0.235186 | 0.057309 | 0.253512 | 0.039387 | 0.260281 | 0.018625 | ... | -0.203892 | 0.027493 | 0.047226 | 0.036891 | -0.043017 | 0.018365 | 0.100931 | -0.013236 | -0.146563 | -0.012573 |
127 rows × 36 columns
We create a DataFrame, called df_shapes_vis, of shape (n_specimens*n_landmarks) x n_dim to visualize the aligned shapes.
df_shapes_vis = df_shapes.copy()
df_shapes_vis.columns = pd.MultiIndex.from_tuples(
[
[int(landmark_idx), dim]
for dim, landmark_idx in [idx.split("_") for idx in df_shapes_vis.columns]
],
names=["coord_id", "dim"],
)
df_shapes_vis.sort_index(axis=1, inplace=True)
df_shapes_vis = df_shapes_vis.swaplevel(0, 1, axis=1).stack(level=1, future_stack=True)
df_shapes_vis
| dim | x | y | |
|---|---|---|---|
| specimen_id | coord_id | ||
| 0 | 0 | -0.488913 | 0.021159 |
| 1 | -0.075651 | 0.083813 | |
| 2 | 0.222655 | 0.081656 | |
| 3 | 0.262641 | 0.041408 | |
| 4 | 0.262701 | 0.021471 | |
| ... | ... | ... | ... |
| 126 | 13 | -0.203892 | 0.027493 |
| 14 | 0.047226 | 0.036891 | |
| 15 | -0.043017 | 0.018365 | |
| 16 | 0.100931 | -0.013236 | |
| 17 | -0.146563 | -0.012573 |
2286 rows × 2 columns
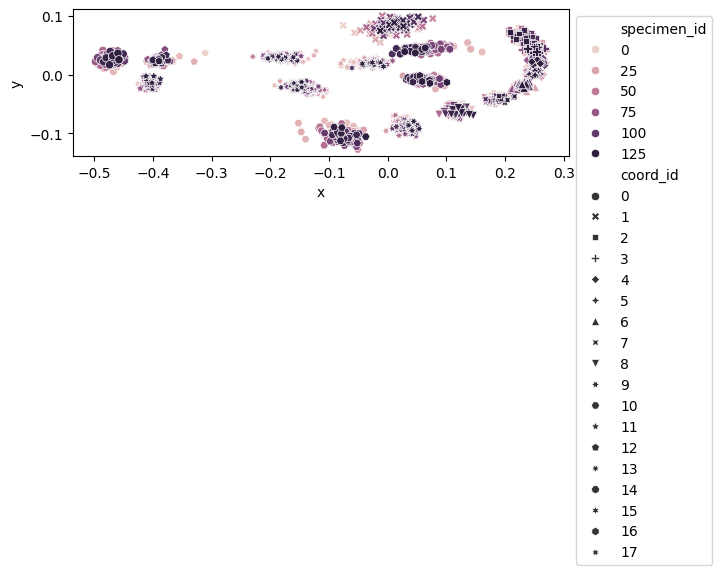
fig, ax = plt.subplots()
sns.scatterplot(
data=df_shapes_vis, x="x", y="y", hue="specimen_id", style="coord_id", ax=ax
)
ax.set_aspect("equal")
ax.legend(loc="upper left", bbox_to_anchor=(1, 1))
<matplotlib.legend.Legend at 0x7fbbf1946690>
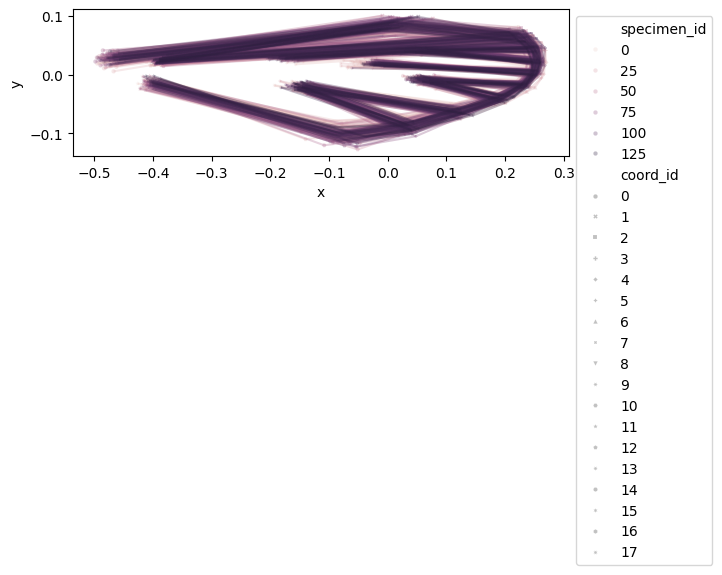
configuration_plot(
df_shapes_vis, links=links, alpha=0.3, hue="specimen_id", style="coord_id"
)
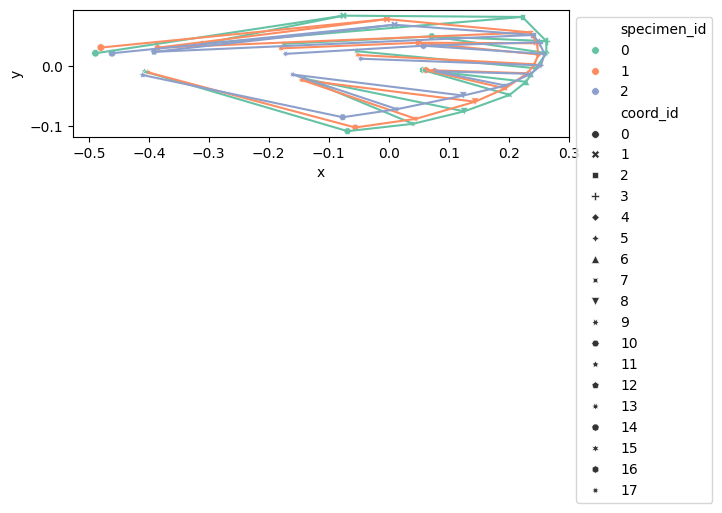
configuration_plot(
df_shapes_vis.loc[0:2],
links=links,
hue="specimen_id",
style="coord_id",
palette="Set2",
s=30,
)
PCA#
pca = PCA(n_components=10).set_output(transform="pandas")
df_pca = pca.fit_transform(df_shapes)
df_pca = df_pca.join(data_landmark_mosquito_wings.meta)
df_pca
| pca0 | pca1 | pca2 | pca3 | pca4 | pca5 | pca6 | pca7 | pca8 | pca9 | genus | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| specimen_id | |||||||||||
| 0 | -0.003856 | -0.093210 | 0.029908 | 0.049589 | -0.007598 | 0.042657 | -0.005370 | 0.010359 | -0.011595 | 0.034991 | AN |
| 1 | -0.029857 | -0.023434 | 0.006114 | -0.017268 | -0.014229 | 0.014271 | 0.013344 | 0.021849 | -0.011211 | 0.005069 | AN |
| 2 | -0.012572 | -0.019728 | -0.034610 | -0.025665 | -0.005116 | -0.017905 | 0.008511 | -0.004312 | 0.001597 | 0.009884 | AN |
| 3 | -0.001652 | -0.049890 | -0.034137 | -0.001013 | 0.019706 | -0.010102 | -0.001692 | -0.010425 | 0.021595 | -0.006472 | AN |
| 4 | 0.026869 | -0.032989 | 0.010228 | -0.051852 | 0.016741 | 0.023749 | 0.015917 | -0.000273 | -0.013100 | 0.012755 | AN |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 122 | -0.007864 | -0.009853 | -0.006369 | -0.026082 | -0.025424 | 0.013786 | -0.009967 | -0.018669 | 0.000513 | -0.005515 | CX |
| 123 | -0.005360 | 0.014451 | -0.014347 | -0.013199 | 0.000102 | 0.001379 | -0.002919 | -0.009755 | -0.020409 | -0.017790 | CX |
| 124 | -0.030056 | 0.000885 | 0.013455 | 0.000542 | -0.008940 | 0.008503 | -0.021390 | -0.014563 | -0.003408 | -0.010083 | CX |
| 125 | -0.048757 | -0.000199 | 0.024147 | -0.029841 | -0.040190 | 0.013335 | 0.000598 | 0.016086 | 0.018985 | 0.004560 | DE |
| 126 | -0.017155 | -0.030018 | -0.001935 | -0.014438 | -0.014540 | -0.035565 | -0.011888 | 0.017324 | -0.009739 | 0.012342 | DE |
127 rows × 11 columns
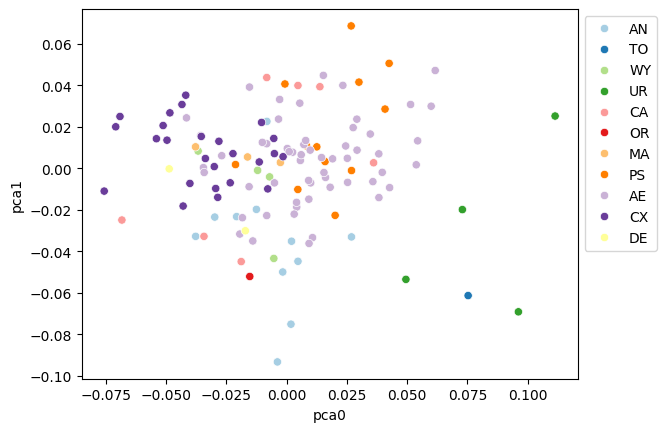
fig, ax = plt.subplots()
sns.scatterplot(data=df_pca, x="pca0", y="pca1", hue="genus", palette="Paired", ax=ax)
ax.legend(loc="upper left", bbox_to_anchor=(1, 1))
<matplotlib.legend.Legend at 0x7fbbf0d42f50>
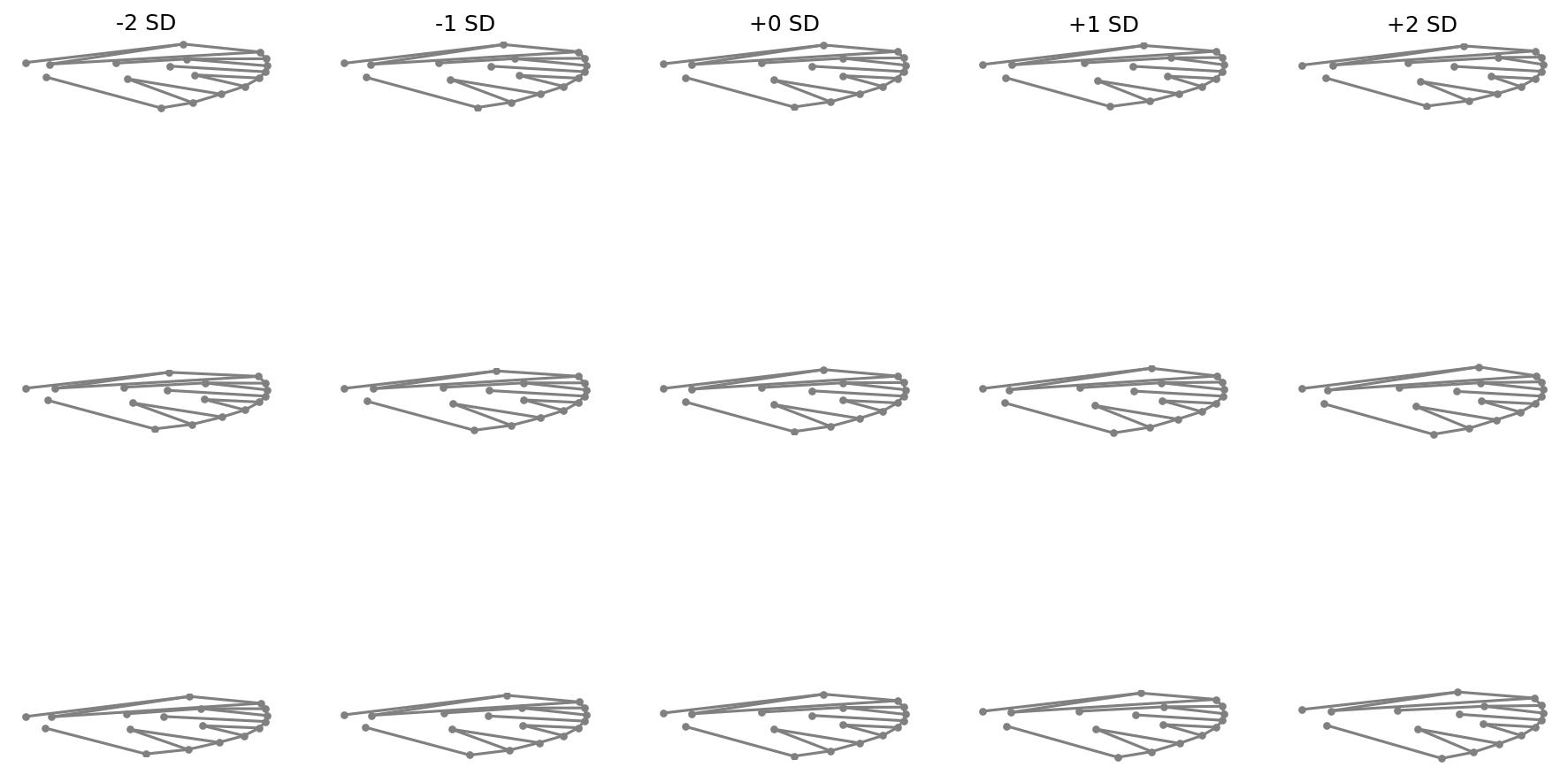
Shape variation along PC axes#
fig = shape_variation_plot(
pca,
n_dim=2,
links=links,
components=(0, 1, 2),
sd_values=(-2, -1, 0, 1, 2),
)
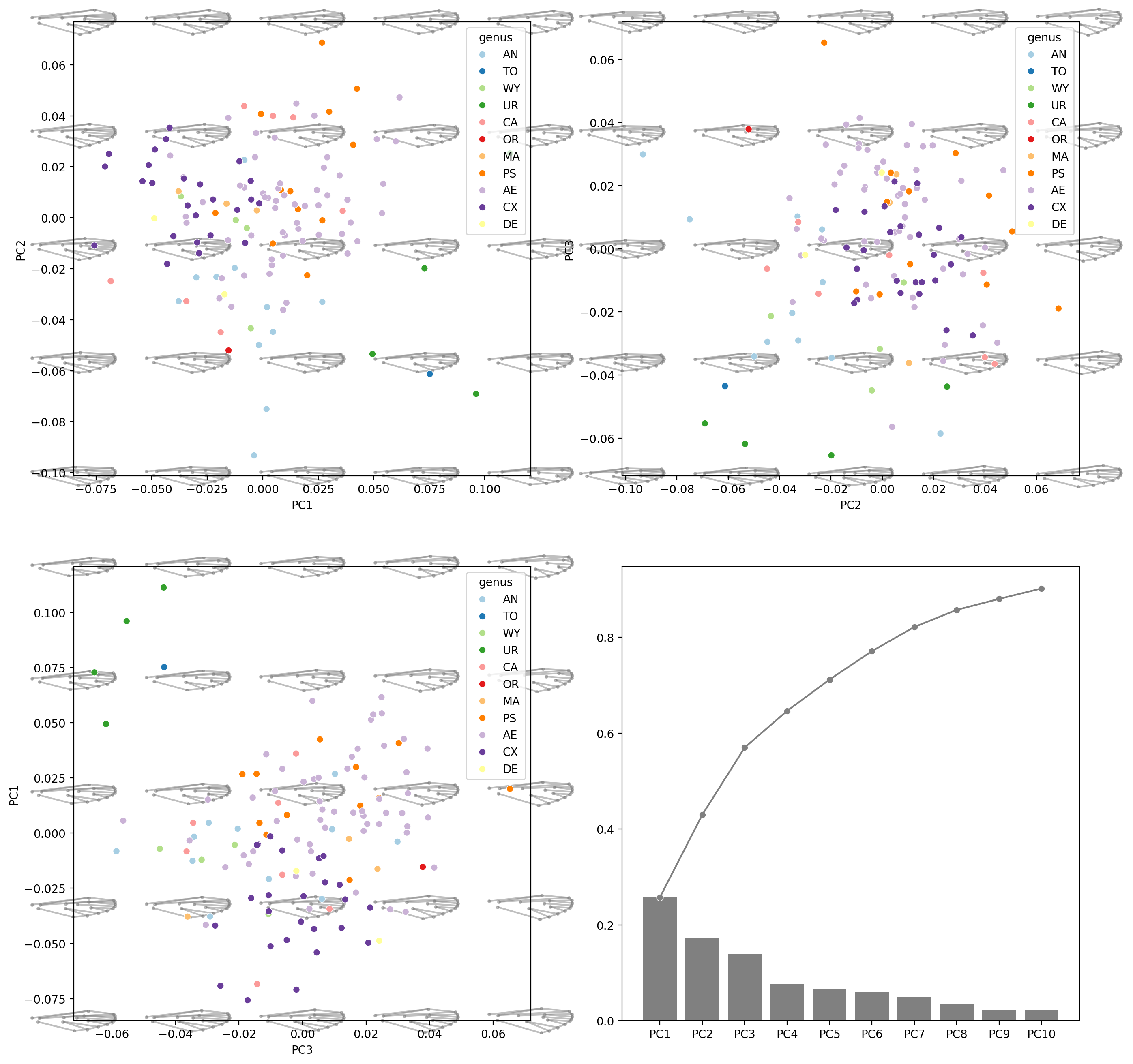
Morphospace#
fig, axes = plt.subplots(2, 2, figsize=(16, 16), dpi=200)
for ax, (i, j) in zip(axes.flat[:3], [(0, 1), (1, 2), (2, 0)]):
morphospace_plot(
data=df_pca,
x=f"pca{i}", y=f"pca{j}", hue="genus",
reducer=pca,
n_dim=2,
components=(i, j),
links=links,
palette="Paired",
n_shapes=5,
shape_color="gray",
shape_scale=1.0,
shape_alpha=0.5,
ax=ax,
s=5,
)
ax.set(xlabel=f"PC{i + 1}", ylabel=f"PC{j + 1}")
explained_variance_ratio_plot(pca, ax=axes[1, 1])
<Axes: >