Semilandmark analysis#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.decomposition import PCA
from ktch.datasets import load_landmark_trilobite_cephala
from ktch.landmark import GeneralizedProcrustesAnalysis, combine_landmarks_and_curves
from ktch.plot import explained_variance_ratio_plot, morphospace_plot
Load trilobite cephalon dataset#
This dataset contains 2D landmark and semilandmark configurations of trilobite cephala (head shields). Each specimen has 16 fixed landmarks and 4 curves with semilandmarks (12, 20, 20, and 20 points respectively).
data = load_landmark_trilobite_cephala()
landmarks = data.landmarks
curves = data.curves
print("landmarks shape:", landmarks.shape)
print("number of curves per specimen:", len(curves[0]))
for i, c in enumerate(curves[0]):
print(f" curve {i}: {c.shape[0]} points")
landmarks shape: (300, 16, 2)
number of curves per specimen: 4
curve 0: 12 points
curve 1: 20 points
curve 2: 20 points
curve 3: 20 points
df_meta = pd.DataFrame(data.meta)
df_meta.head()
| genus | species | ref_pic | locality | formation | min_age | max_age | format | has_scale | |
|---|---|---|---|---|---|---|---|---|---|
| 1020_Liu_1977 | Hunanolenus | Hunanolenus genalatus | Liu 1977 | Hunan | Huaqiao | Paibian | Paibian | xml | True |
| 1023_Liu_1977 | Huangshiaspis | Huangshiaspis transversus | Liu 1977 | Fenghuang area | Tingziguan | Jiangshanian | Jiangshanian | xml | True |
| AMF27749 | Weania | Weania semicircularis | Vanderlaan y Ebach 2015 | Glen William Road | Flagstaff | Middle Visean | Upper Visean | xml | True |
| AMNH_28896 | Eldredgeops | Eldredgeops raNAmilleri | Burton and Eldredge 1974 | NaN | NaN | Eifelian | NaN | tps | True |
| AMNH_28902 | Viaphacops | Viaphacops stummi | Eldredge 1973 | NaN | NaN | Eifelian | NaN | tps | True |
Visualize raw data#
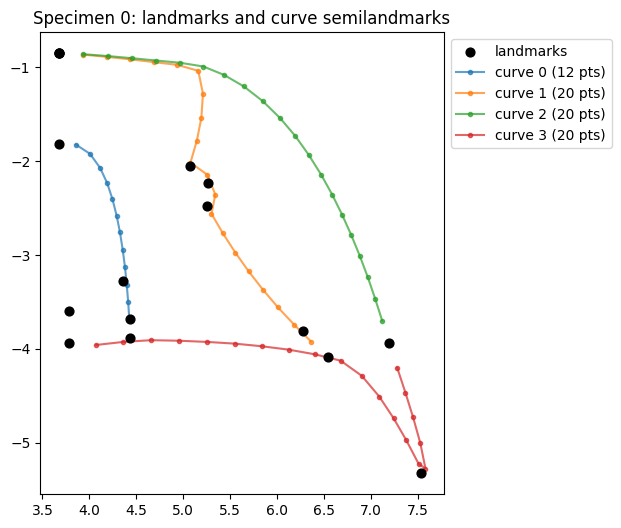
Plot a single specimen, distinguishing fixed landmarks from curve semilandmarks.
specimen_idx = 0
fig, ax = plt.subplots(figsize=(6, 6))
# Fixed landmarks
lm = landmarks[specimen_idx]
ax.scatter(lm[:, 0], lm[:, 1], c="black", s=40, zorder=3, label="landmarks")
# Curve semilandmarks
curve_colors = ["tab:blue", "tab:orange", "tab:green", "tab:red"]
for i, curve in enumerate(curves[specimen_idx]):
ax.plot(curve[:, 0], curve[:, 1], "o-", color=curve_colors[i], markersize=3, alpha=0.7, label=f"curve {i} ({curve.shape[0]} pts)")
ax.set_aspect("equal")
ax.legend(loc="upper left", bbox_to_anchor=(1, 1))
ax.set_title("Specimen 0: landmarks and curve semilandmarks")
Text(0.5, 1.0, 'Specimen 0: landmarks and curve semilandmarks')
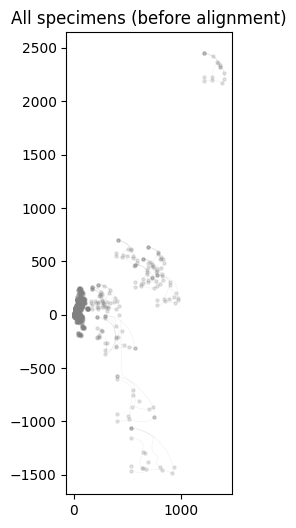
Plot all specimens overlaid before alignment.
fig, ax = plt.subplots(figsize=(6, 6))
for i in range(landmarks.shape[0]):
lm = landmarks[i]
ax.scatter(lm[:, 0], lm[:, 1], c="gray", s=5, alpha=0.2)
for curve in curves[i]:
ax.plot(curve[:, 0], curve[:, 1], c="gray", alpha=0.1, linewidth=0.5)
ax.set_aspect("equal")
ax.set_title("All specimens (before alignment)")
Text(0.5, 1.0, 'All specimens (before alignment)')
Combine landmarks and curves#
combine_landmarks_and_curves() merges the fixed landmarks and curve semilandmarks
into a single configuration array and generates the slider matrix that defines the sliding topology.
The curve_landmarks parameter specifies which landmarks anchor each curve.
Each curve’s semilandmarks slide along the tangent direction, with the
specified landmarks serving as fixed endpoints.
combined, slider_matrix, curve_info = combine_landmarks_and_curves(
landmarks,
curves,
curve_landmarks=data.curve_landmarks,
)
print("combined shape:", combined.shape)
print("slider_matrix shape:", slider_matrix.shape)
print("curve_info:", curve_info)
combined shape: (300, 88, 2)
slider_matrix shape: (72, 3)
curve_info: {'n_landmarks': 16, 'n_curve_points': 72, 'curve_offsets': [16, 28, 48, 68], 'curve_lengths': [12, 20, 20, 20]}
The slider matrix has shape (n_sliders, 3) where each row is [before_index, slider_index, after_index].
All semilandmarks are included in the slider matrix. For curve endpoints,
the before/after neighbor is the anchoring landmark.
print("First 5 rows of slider_matrix:")
print(slider_matrix[:5])
First 5 rows of slider_matrix:
[[ 1 16 17]
[16 17 18]
[17 18 19]
[18 19 20]
[19 20 21]]
Reshape the combined array to 2D (n_specimens, n_points * n_dim) for GPA input.
Specimens containing NaN values are excluded.
X = combined.reshape(combined.shape[0], -1)
nan_mask = np.any(np.isnan(X), axis=1)
print(f"Specimens with NaN: {nan_mask.sum()} / {X.shape[0]}")
X_clean = X[~nan_mask]
df_meta_clean = df_meta[~nan_mask].reset_index(drop=True)
print(f"Specimens used for analysis: {X_clean.shape[0]}")
Specimens with NaN: 15 / 300
Specimens used for analysis: 285
GPA with semilandmark sliding#
Pass the curves parameter to GeneralizedProcrustesAnalysis to enable semilandmark sliding
during Procrustes superimposition. Semilandmarks slide along their curve tangent directions
to minimize bending energy.
The tol parameter controls convergence tolerance for the iterative GPA algorithm.
gpa = GeneralizedProcrustesAnalysis(n_dim=2, curves=slider_matrix, tol=1e-3)
shapes = gpa.fit_transform(X_clean)
print("shapes shape:", shapes.shape)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 70 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 69 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
/home/runner/work/ktch/ktch/ktch/landmark/_procrustes_analysis.py:432: UserWarning: Rank-deficient sliding system: rank 71 < 72. Tangent directions may be degenerate (zero-length or collinear).
x_slid = slide_func(x_rot, mu, curves)
shapes shape: (285, 176)
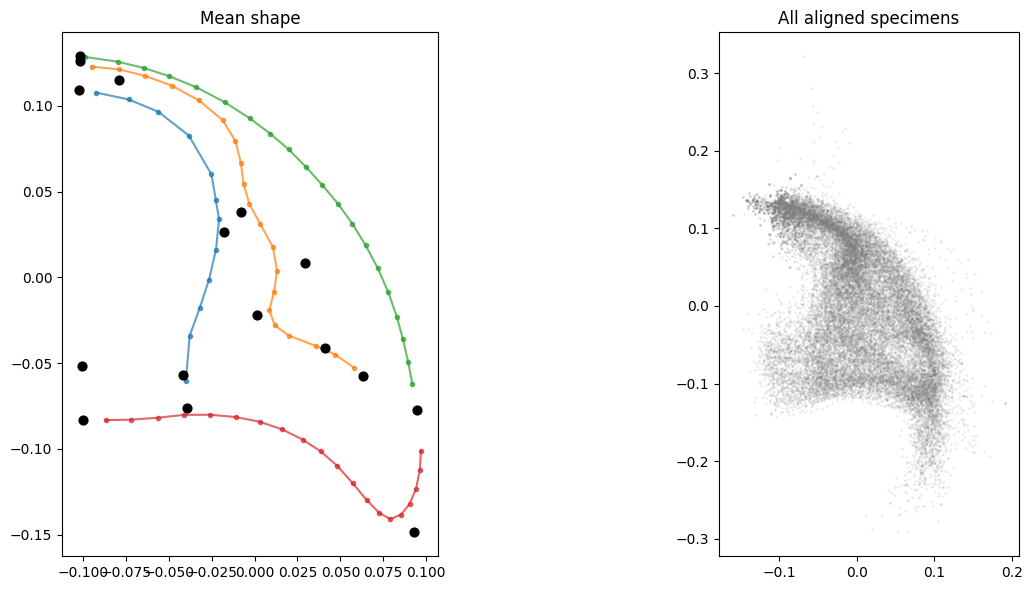
Visualize the aligned shapes.
n_points = combined.shape[1]
n_dim = 2
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
# Mean shape
mean_shape = shapes.mean(axis=0).reshape(n_points, n_dim)
axes[0].scatter(mean_shape[:16, 0], mean_shape[:16, 1], c="black", s=40, zorder=3)
offset = 16
for i, length in enumerate(curve_info["curve_lengths"]):
start = offset
end = offset + length
axes[0].plot(
mean_shape[start:end, 0], mean_shape[start:end, 1],
"o-", color=curve_colors[i], markersize=3, alpha=0.7,
)
offset = end
axes[0].set_aspect("equal")
axes[0].set_title("Mean shape")
# All aligned specimens
for i in range(shapes.shape[0]):
specimen = shapes[i].reshape(n_points, n_dim)
axes[1].scatter(specimen[:, 0], specimen[:, 1], c="gray", s=1, alpha=0.1)
axes[1].set_aspect("equal")
axes[1].set_title("All aligned specimens")
plt.tight_layout()
PCA#
pca = PCA(n_components=10)
pc_scores = pca.fit_transform(shapes)
df_pca = pd.DataFrame(
pc_scores,
columns=[f"PC{i+1}" for i in range(pc_scores.shape[1])],
)
df_pca = df_pca.join(df_meta_clean[["genus"]])
df_pca.head()
| PC1 | PC2 | PC3 | PC4 | PC5 | PC6 | PC7 | PC8 | PC9 | PC10 | genus | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | -0.125228 | -0.162647 | 0.004561 | 0.050726 | 0.019523 | 0.050510 | -0.016427 | 0.043711 | 0.002174 | -0.025951 | Hunanolenus |
| 1 | -0.041252 | -0.236899 | 0.140655 | 0.112496 | 0.036621 | 0.002261 | 0.001618 | 0.024187 | -0.056088 | 0.014121 | Huangshiaspis |
| 2 | 0.009868 | -0.111546 | -0.027055 | -0.132076 | 0.009976 | -0.019524 | 0.101439 | -0.009635 | 0.039078 | -0.067129 | Weania |
| 3 | 0.142255 | 0.099790 | 0.069591 | -0.034382 | -0.008973 | 0.026952 | 0.003910 | 0.059150 | -0.012461 | 0.021385 | Eldredgeops |
| 4 | 0.088966 | 0.111502 | 0.055935 | -0.030005 | 0.049822 | 0.041755 | 0.008725 | 0.068558 | -0.035017 | 0.035339 | Viaphacops |
fig, ax = plt.subplots()
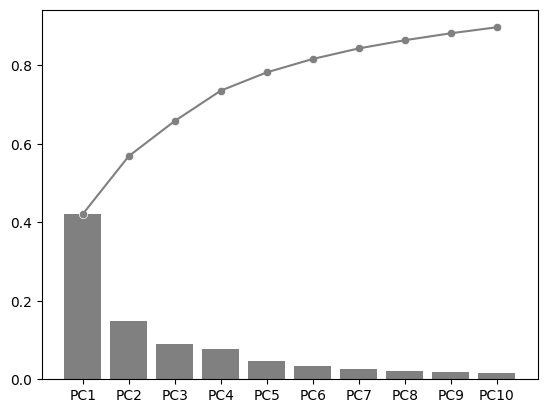
explained_variance_ratio_plot(pca, ax=ax, verbose=True)
PC1 PC2 PC3 PC4 PC5 PC6 PC7 PC8 PC9 PC10
Explained Variance Ratio (EVR) 0.4207 0.1478 0.0897 0.0773 0.0466 0.0342 0.0269 0.0209 0.0177 0.0153
Cumsum of EVR 0.4207 0.5685 0.6582 0.7355 0.7821 0.8163 0.8432 0.8641 0.8818 0.8971
<Axes: >
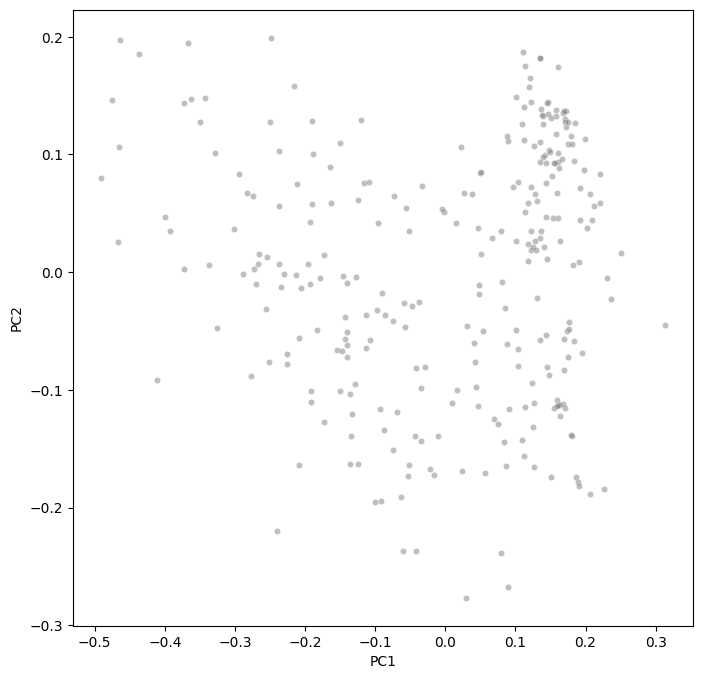
Morphospace visualization#
fig, ax = plt.subplots(figsize=(8, 8))
sns.scatterplot(
data=df_pca,
x="PC1",
y="PC2",
ax=ax,
c="gray",
alpha=0.5,
s=20,
)
ax.set(xlabel="PC1", ylabel="PC2")
[Text(0.5, 0, 'PC1'), Text(0, 0.5, 'PC2')]
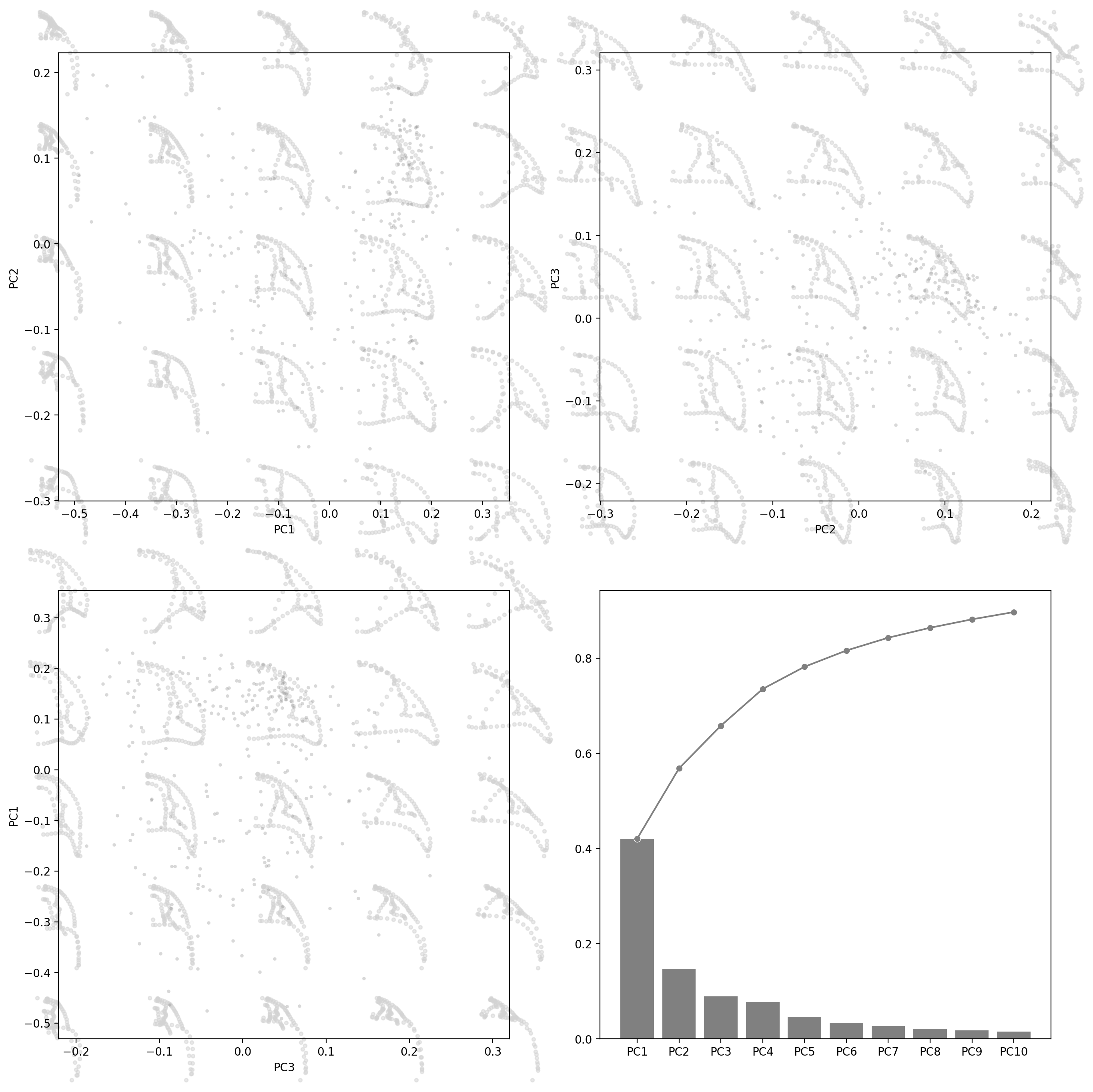
fig, axes = plt.subplots(2, 2, figsize=(16, 16), dpi=200)
for ax, (i, j) in zip(axes.flat[:3], [(0, 1), (1, 2), (2, 0)]):
morphospace_plot(
data=df_pca,
x=f"PC{i + 1}", y=f"PC{j + 1}",
reducer=pca,
n_dim=2,
shape_type="landmarks_2d",
components=(i, j),
n_shapes=5,
shape_alpha=0.5,
ax=ax,
scatter_kw=dict(c="gray", alpha=0.3, s=10),
)
explained_variance_ratio_plot(pca, ax=axes[1, 1], verbose=True)
PC1 PC2 PC3 PC4 PC5 PC6 PC7 PC8 PC9 PC10
Explained Variance Ratio (EVR) 0.4207 0.1478 0.0897 0.0773 0.0466 0.0342 0.0269 0.0209 0.0177 0.0153
Cumsum of EVR 0.4207 0.5685 0.6582 0.7355 0.7821 0.8163 0.8432 0.8641 0.8818 0.8971
<Axes: >