2D Outline Extraction from Images#
This tutorial shows how to extract outlines from images based on conventional image processing methods for use with ktch’s Elliptic Fourier Analysis (EFA).
Prerequisites#
# !pip install ktch[data] opencv-python # Uncomment if needed
Step 1: Load Images#
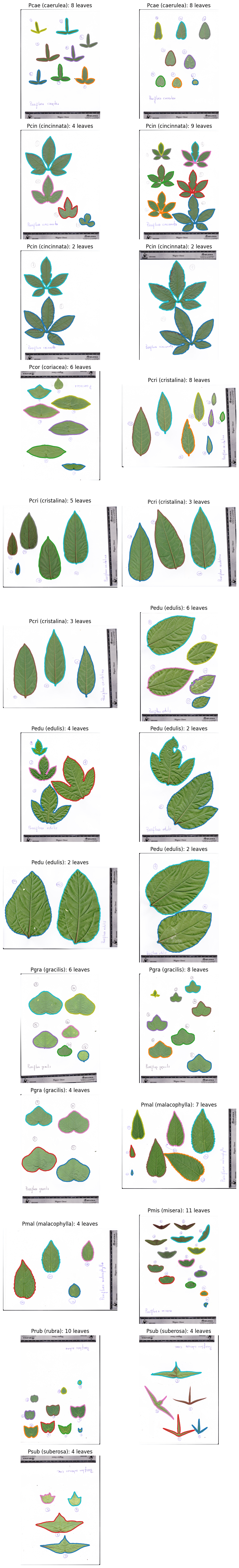
We use the Passiflora leaf scan dataset bundled with ktch: 25 flatbed scan images
of 10 Passiflora species spanning simple elliptical to deeply lobed leaf forms.
Each image contains multiple leaves from one plant individual, arranged from tip
(youngest) to base (oldest). This dataset is a subset of the Passiflora leaf scan data from
Chitwood & Otoni (2017)
(data: Chitwood & Otoni, 2016).
See data.DESCR for full details.
import numpy as np
import cv2 as cv
import matplotlib.pyplot as plt
from ktch.datasets import load_image_passiflora_leaves
data = load_image_passiflora_leaves(as_frame=True)
print(f"Number of images: {len(data.images)}")
print(f"Species: {data.meta['species'].nunique()}")
print()
print(data.meta[["abbreviation", "species"]].value_counts().sort_index())
Number of images: 25
Species: 10
abbreviation species
Pcae caerulea 2
Pcin cincinnata 4
Pcor coriacea 1
Pcri cristalina 4
Pedu edulis 5
Pgra gracilis 3
Pmal malacophylla 2
Pmis misera 1
Prub rubra 1
Psub suberosa 2
Name: count, dtype: int64
labels = [
f"{row.abbreviation} ({row.species})"
for _, row in data.meta.iterrows()
]
plot_images(data.images, labels=labels)
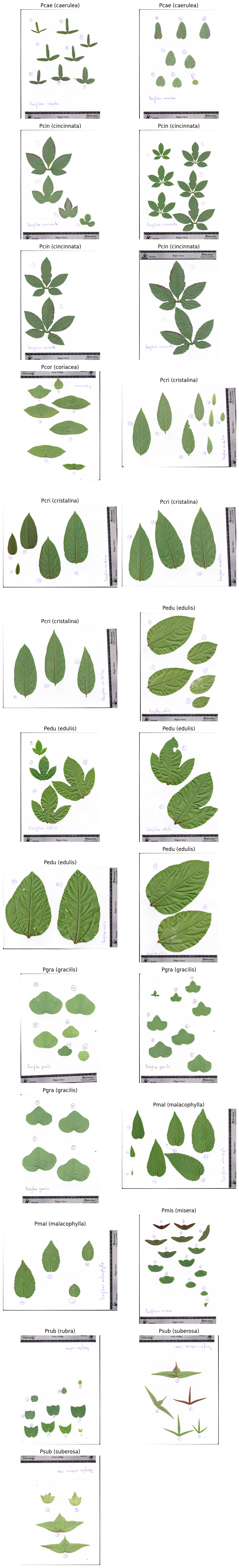

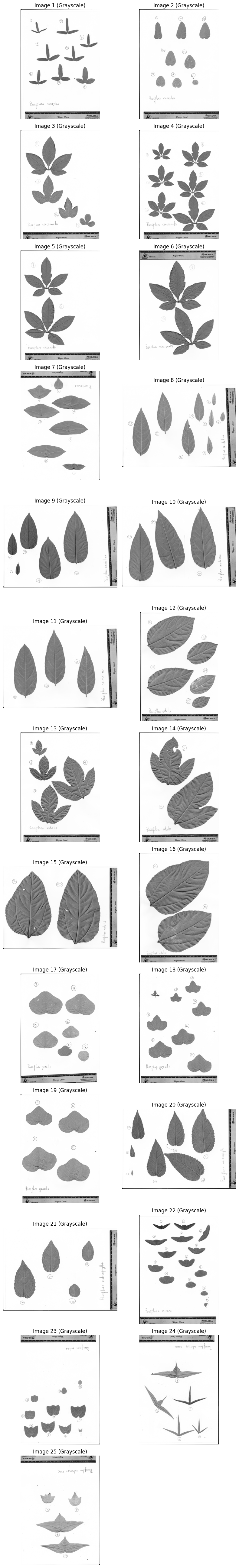
Step 2: Convert to Grayscale#
images_gray = [cv.cvtColor(img, cv.COLOR_RGB2GRAY) for img in data.images]
plot_images(images_gray, "Grayscale", cmap="gray")
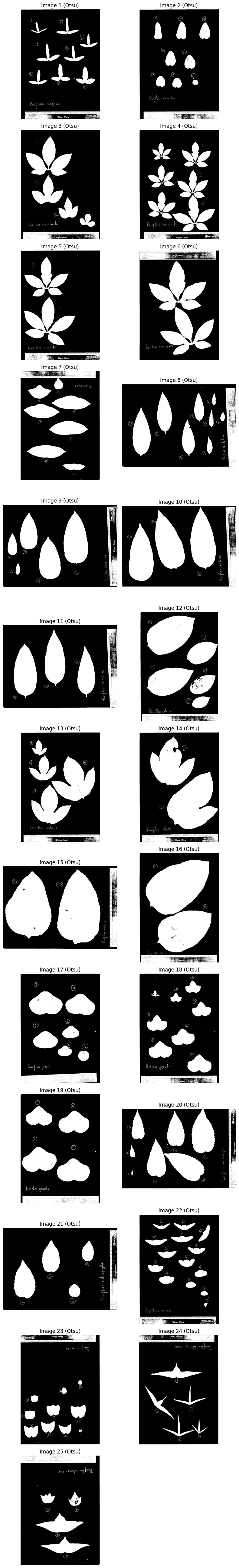
Step 3: Binarize the Image#
We combine two binarization methods with OR to capture leaves that are either darker than the background or green in color.
Otsu’s thresholding#
Automatically finds a threshold based on brightness histogram.
images_binary_otsu = []
for img_gray in images_gray:
_, binary_otsu = cv.threshold(
img_gray, 0, 255, cv.THRESH_BINARY_INV + cv.THRESH_OTSU
)
images_binary_otsu.append(binary_otsu)
plot_images(images_binary_otsu, "Otsu", cmap="gray")
Green mask in HSV#
Targets green hues with sufficient saturation and value.
hue_min, hue_max = 30, 90
sat_min, val_min = 40, 40
images_binary_green = []
for img_rgb in data.images:
img_hsv = cv.cvtColor(img_rgb, cv.COLOR_RGB2HSV)
lower_green = np.array([hue_min, sat_min, val_min])
upper_green = np.array([hue_max, 255, 255])
binary_green = cv.inRange(img_hsv, lower_green, upper_green)
images_binary_green.append(binary_green)
plot_images(images_binary_green, "Green Mask", cmap="gray")
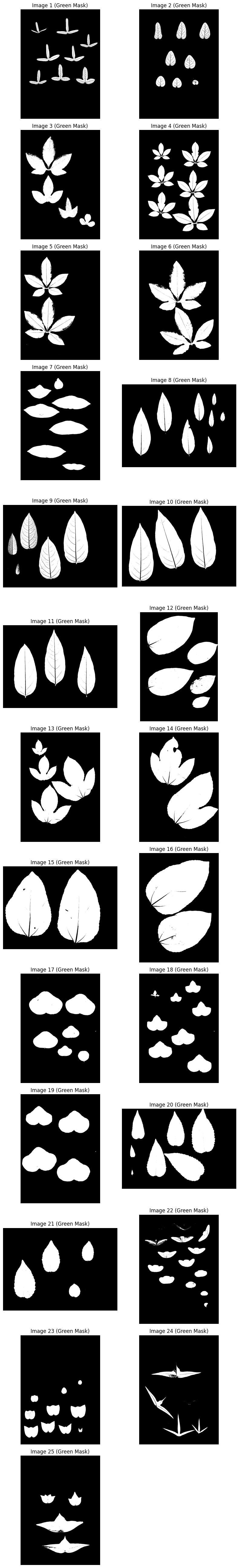

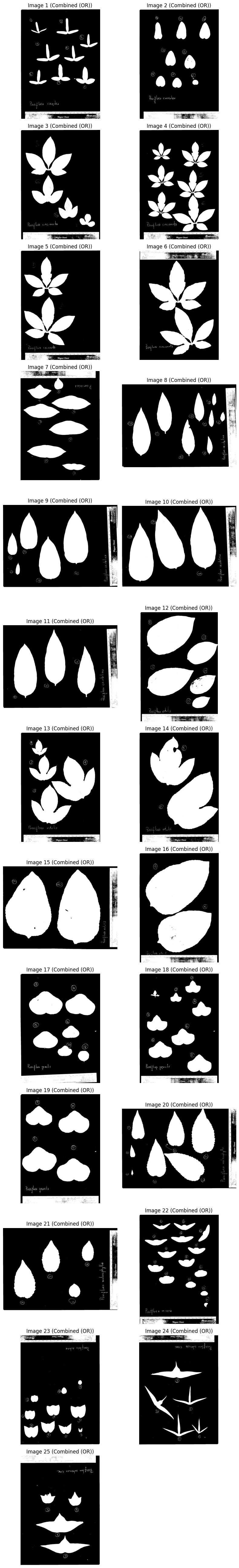
Combine with OR#
images_binary = [
cv.bitwise_or(otsu, green)
for otsu, green in zip(images_binary_otsu, images_binary_green)
]
plot_images(images_binary, "Combined (OR)", cmap="gray")
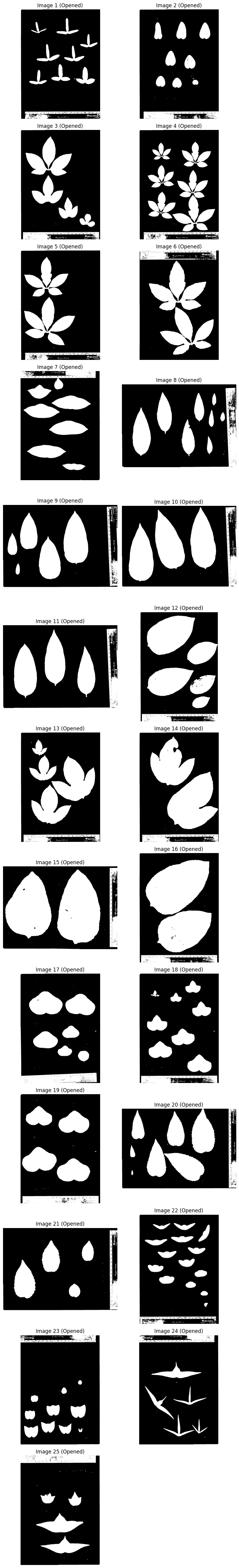
Step 4: Morphological Opening#
Remove small noise with morphological opening.
kernel_size = 5
kernel = cv.getStructuringElement(cv.MORPH_ELLIPSE, (kernel_size, kernel_size))
images_opened = [
cv.morphologyEx(img_binary, cv.MORPH_OPEN, kernel)
for img_binary in images_binary
]
plot_images(images_opened, "Opened", cmap="gray")
Step 5: Extract Outlines#
Extract contours and filter by area and color.
Warning
This simple pipeline does not work perfectly for all images.
For example, deeply lobed species like P. cincinnata may have individual leaflets detected as separate leaves, and pale-colored leaves in P. edulis scans may be partially merged with the background, producing jagged outlines.
These cases require species-specific tuning or more advanced segmentation methods, but are left as-is here to illustrate typical failure cases.
Color filter function#
def is_green_region(img_rgb, contour, hue_range=(30, 90), min_green_ratio=0.5):
"""Check if the contour region contains enough green pixels."""
mask = np.zeros(img_rgb.shape[:2], dtype=np.uint8)
cv.drawContours(mask, [contour], 0, 255, -1)
region_pixels = img_rgb[mask == 255]
if len(region_pixels) == 0:
return False
region_rgb = region_pixels.reshape(-1, 1, 3)
region_hsv = cv.cvtColor(region_rgb, cv.COLOR_RGB2HSV).reshape(-1, 3)
h, s, v = region_hsv[:, 0], region_hsv[:, 1], region_hsv[:, 2]
is_green = (h >= hue_range[0]) & (h <= hue_range[1]) & (s >= 40) & (v >= 40)
return np.sum(is_green) / len(region_pixels) >= min_green_ratio
Filter parameters#
min_area = 50
hue_range = (30, 90)
min_green_ratio = 0.5
Extract and filter contours#
all_outlines = []
outlines_per_image = []
for idx, img_opened in enumerate(images_opened):
img_rgb = data.images[idx]
contours, _ = cv.findContours(
img_opened, cv.RETR_EXTERNAL, cv.CHAIN_APPROX_NONE
)
contours_filtered = [
c for c in contours
if cv.contourArea(c) >= min_area
and is_green_region(img_rgb, c, hue_range, min_green_ratio)
]
outlines = [c[:, 0, :].astype(np.float64) for c in contours_filtered]
outlines_per_image.append(outlines)
all_outlines.extend(outlines)
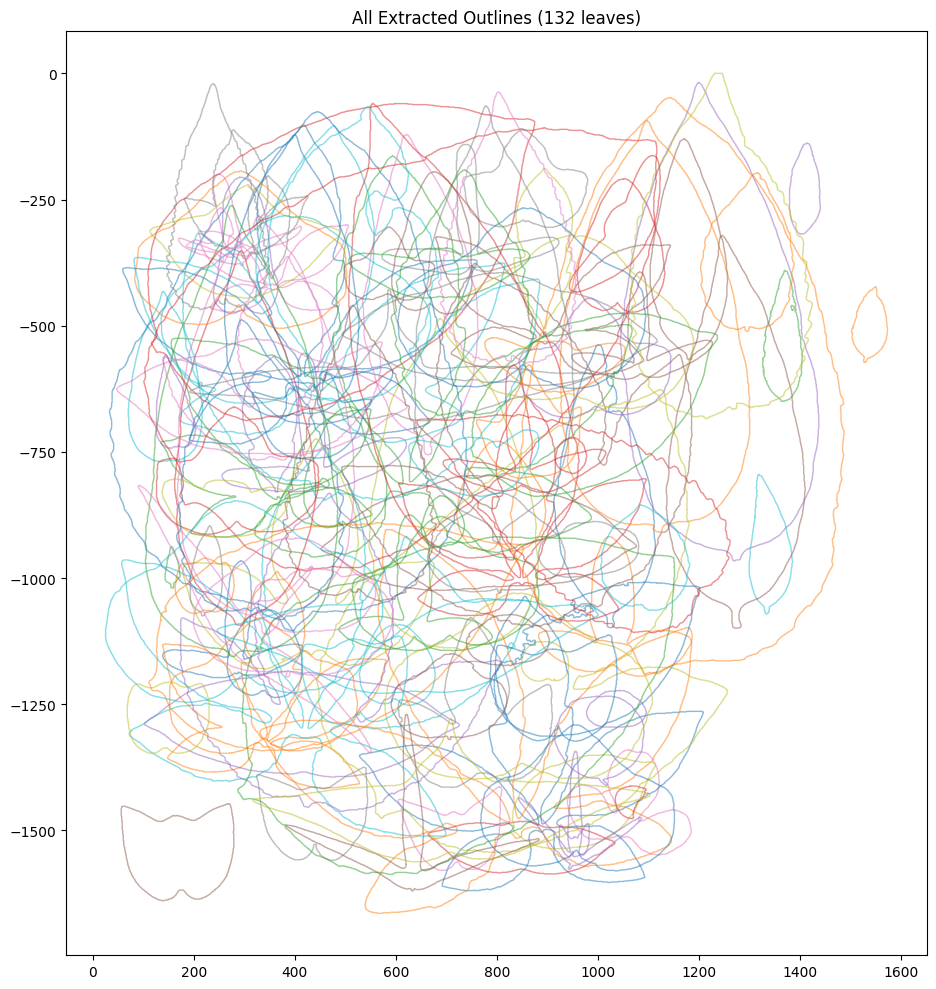
print(f"Total: {len(all_outlines)} outlines")
Total: 132 outlines
plot_images(data.images, outlines_per_image=outlines_per_image, labels=labels)
if len(all_outlines) > 0:
fig, ax = plt.subplots(figsize=(10, 10))
for outline in all_outlines:
ax.plot(outline[:, 0], -outline[:, 1], alpha=0.5, linewidth=1)
ax.set_aspect("equal")
ax.set_title(f"All Extracted Outlines ({len(all_outlines)} leaves)")
plt.tight_layout()
else:
print("No outlines to plot")
Step 6: Verify Output Format#
The output format is compatible with ktch’s EllipticFourierAnalysis.
print(f"Type: {type(all_outlines)}")
print(f"Number of specimens: {len(all_outlines)}")
if len(all_outlines) == 0:
print("\nWarning: No outlines detected. Try adjusting parameters:")
print(" - Decrease min_area")
print(" - Adjust hue_range for your object color")
print(" - Decrease min_green_ratio")
else:
print(f"Shape of first outline: {all_outlines[0].shape}")
Type: <class 'list'>
Number of specimens: 132
Shape of first outline: (935, 2)
from ktch.harmonic import EllipticFourierAnalysis
if len(all_outlines) > 0:
efa = EllipticFourierAnalysis(n_harmonics=20)
coef = efa.fit_transform(all_outlines)
print(f"EFA output shape: {coef.shape}")
else:
print("Skipping EFA: no outlines to process")
EFA output shape: (132, 84)
Summary#
This tutorial demonstrated a complete pipeline for extracting outlines from images: grayscale conversion, binarization (Otsu + green mask), morphological opening, and contour filtering. The extracted outlines are ready for EFA analysis with ktch.
Next Steps#
Elliptic Fourier Analysis - EFA with PCA visualization
Load CHC Files - Save/load outline data
Harmonic-based Morphometrics - Theory behind harmonic analysis
Troubleshooting#
Too few contours#
Decrease
min_areato include smaller regionsWiden
hue_rangeor lowermin_green_ratiofor less strict color filteringDecrease
kernel_sizeto preserve fine details
Too many contours (noise)#
Increase
min_areato filter out small artifactsIncrease
kernel_sizefor stronger noise removalRaise
min_green_ratiofor stricter color filtering
Inconsistent orientation#
EFA results depend on outline direction (clockwise vs counter-clockwise). Compute signed area and reverse outlines if negative to standardize.
Holes or gaps in binary image#
Apply morphological closing after opening:
cv.morphologyEx(img, cv.MORPH_CLOSE, kernel)Adjust binarization thresholds or HSV ranges